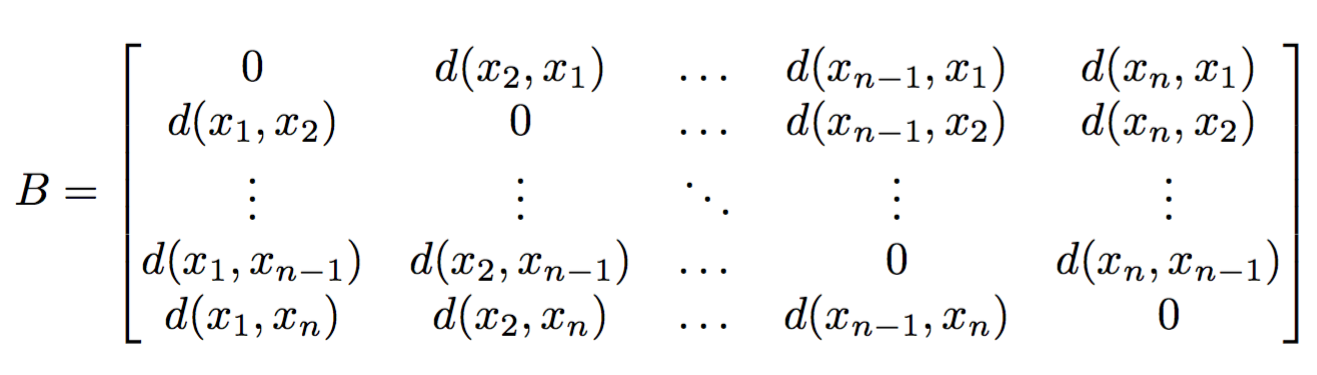Outline¶
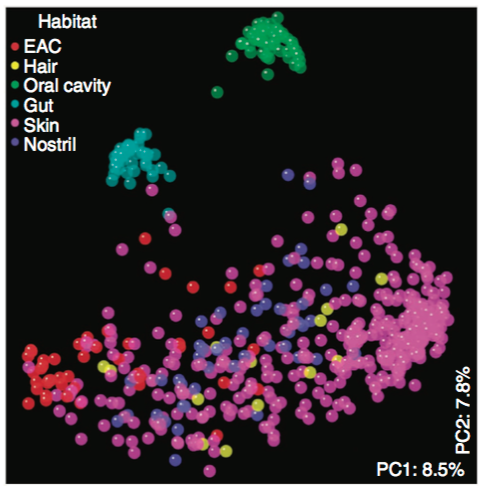
Background (why $\beta$-diversity).What is Emperor.
How can we use Emperor.
Analyzing a use case.
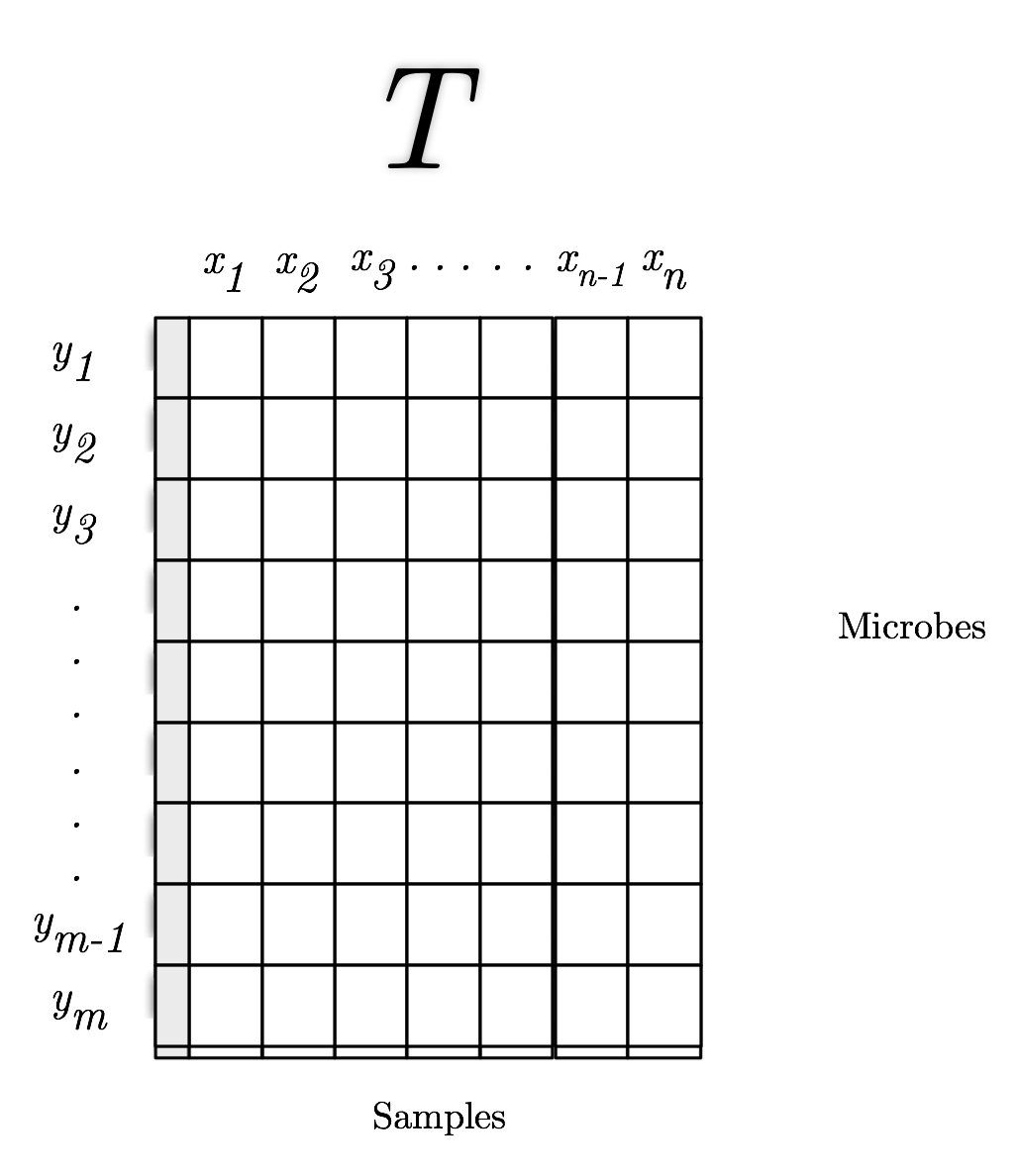
What is Emperor?¶
A Python 2/3 package that powers a JavaScript UI.
Originated in the context of Quantitative Insights Into Microbial Ecology http://2.qiime.org
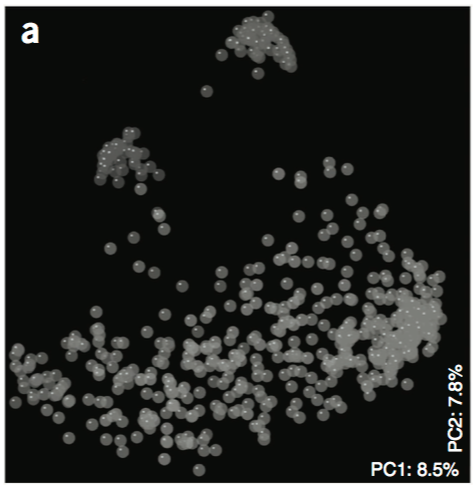
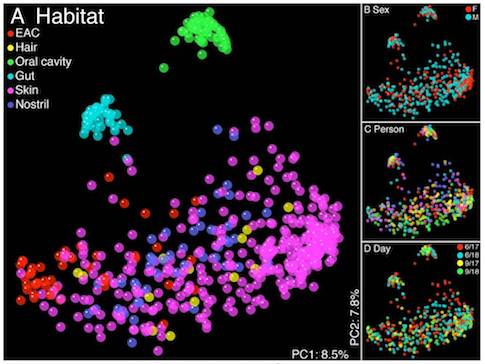
Scatter plot viewer.
- Visualize
- Interact
- Share
Other applications¶
- Qiita

- American Gut (participant's results).
- Illumina's BaseSpace processing results for the QIIME app.
- Metabolomic analysis, IPython notebook ...
... kinda
Nowadays¶
- Python API.
- Python 2 and 3.
- Integration with scikit-bio.
- Jupyter integration.
- Pandas 🐼 Integration.
- JavaScript API.
- Command Line Interface (powered by QIIME 2).
Outline¶
Background (why $\beta$-diversity).What is Emperor.How can we use Emperor.
Analyzing a use case.
Python¶
1 main class
Emperor, depends onscikit-bioandpandas.Format Python data into JSON and display it using JavaScript.
from emperor import Emperor
Emperor?
🐼s integration - experimental¶
import pandas as pd
df = pd.read_csv('./tips.csv')
df
| total_bill | tip | sex | smoker | day | time | size | |
|---|---|---|---|---|---|---|---|
| 0 | 16.99 | 1.01 | Female | No | Sun | Dinner | 2 |
| 1 | 10.34 | 1.66 | Male | No | Sun | Dinner | 3 |
| 2 | 21.01 | 3.50 | Male | No | Sun | Dinner | 3 |
| 3 | 23.68 | 3.31 | Male | No | Sun | Dinner | 2 |
| 4 | 24.59 | 3.61 | Female | No | Sun | Dinner | 4 |
| 5 | 25.29 | 4.71 | Male | No | Sun | Dinner | 4 |
| 6 | 8.77 | 2.00 | Male | No | Sun | Dinner | 2 |
| 7 | 26.88 | 3.12 | Male | No | Sun | Dinner | 4 |
| 8 | 15.04 | 1.96 | Male | No | Sun | Dinner | 2 |
| 9 | 14.78 | 3.23 | Male | No | Sun | Dinner | 2 |
| 10 | 10.27 | 1.71 | Male | No | Sun | Dinner | 2 |
| 11 | 35.26 | 5.00 | Female | No | Sun | Dinner | 4 |
| 12 | 15.42 | 1.57 | Male | No | Sun | Dinner | 2 |
| 13 | 18.43 | 3.00 | Male | No | Sun | Dinner | 4 |
| 14 | 14.83 | 3.02 | Female | No | Sun | Dinner | 2 |
| 15 | 21.58 | 3.92 | Male | No | Sun | Dinner | 2 |
| 16 | 10.33 | 1.67 | Female | No | Sun | Dinner | 3 |
| 17 | 16.29 | 3.71 | Male | No | Sun | Dinner | 3 |
| 18 | 16.97 | 3.50 | Female | No | Sun | Dinner | 3 |
| 19 | 20.65 | 3.35 | Male | No | Sat | Dinner | 3 |
| 20 | 17.92 | 4.08 | Male | No | Sat | Dinner | 2 |
| 21 | 20.29 | 2.75 | Female | No | Sat | Dinner | 2 |
| 22 | 15.77 | 2.23 | Female | No | Sat | Dinner | 2 |
| 23 | 39.42 | 7.58 | Male | No | Sat | Dinner | 4 |
| 24 | 19.82 | 3.18 | Male | No | Sat | Dinner | 2 |
| 25 | 17.81 | 2.34 | Male | No | Sat | Dinner | 4 |
| 26 | 13.37 | 2.00 | Male | No | Sat | Dinner | 2 |
| 27 | 12.69 | 2.00 | Male | No | Sat | Dinner | 2 |
| 28 | 21.70 | 4.30 | Male | No | Sat | Dinner | 2 |
| 29 | 19.65 | 3.00 | Female | No | Sat | Dinner | 2 |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 214 | 28.17 | 6.50 | Female | Yes | Sat | Dinner | 3 |
| 215 | 12.90 | 1.10 | Female | Yes | Sat | Dinner | 2 |
| 216 | 28.15 | 3.00 | Male | Yes | Sat | Dinner | 5 |
| 217 | 11.59 | 1.50 | Male | Yes | Sat | Dinner | 2 |
| 218 | 7.74 | 1.44 | Male | Yes | Sat | Dinner | 2 |
| 219 | 30.14 | 3.09 | Female | Yes | Sat | Dinner | 4 |
| 220 | 12.16 | 2.20 | Male | Yes | Fri | Lunch | 2 |
| 221 | 13.42 | 3.48 | Female | Yes | Fri | Lunch | 2 |
| 222 | 8.58 | 1.92 | Male | Yes | Fri | Lunch | 1 |
| 223 | 15.98 | 3.00 | Female | No | Fri | Lunch | 3 |
| 224 | 13.42 | 1.58 | Male | Yes | Fri | Lunch | 2 |
| 225 | 16.27 | 2.50 | Female | Yes | Fri | Lunch | 2 |
| 226 | 10.09 | 2.00 | Female | Yes | Fri | Lunch | 2 |
| 227 | 20.45 | 3.00 | Male | No | Sat | Dinner | 4 |
| 228 | 13.28 | 2.72 | Male | No | Sat | Dinner | 2 |
| 229 | 22.12 | 2.88 | Female | Yes | Sat | Dinner | 2 |
| 230 | 24.01 | 2.00 | Male | Yes | Sat | Dinner | 4 |
| 231 | 15.69 | 3.00 | Male | Yes | Sat | Dinner | 3 |
| 232 | 11.61 | 3.39 | Male | No | Sat | Dinner | 2 |
| 233 | 10.77 | 1.47 | Male | No | Sat | Dinner | 2 |
| 234 | 15.53 | 3.00 | Male | Yes | Sat | Dinner | 2 |
| 235 | 10.07 | 1.25 | Male | No | Sat | Dinner | 2 |
| 236 | 12.60 | 1.00 | Male | Yes | Sat | Dinner | 2 |
| 237 | 32.83 | 1.17 | Male | Yes | Sat | Dinner | 2 |
| 238 | 35.83 | 4.67 | Female | No | Sat | Dinner | 3 |
| 239 | 29.03 | 5.92 | Male | No | Sat | Dinner | 3 |
| 240 | 27.18 | 2.00 | Female | Yes | Sat | Dinner | 2 |
| 241 | 22.67 | 2.00 | Male | Yes | Sat | Dinner | 2 |
| 242 | 17.82 | 1.75 | Male | No | Sat | Dinner | 2 |
| 243 | 18.78 | 3.00 | Female | No | Thur | Dinner | 2 |
244 rows × 7 columns
🐼s integration - experimental¶
- Coordinates are inferred from the numerical data, see additional
x,yandzparameters.
from emperor import scatterplot
scatterplot(df, remote=False)

<emperor.core.Emperor at 0x1176b74e0>