Pathway pYPK0_SsXYL1_SsXYL2¶
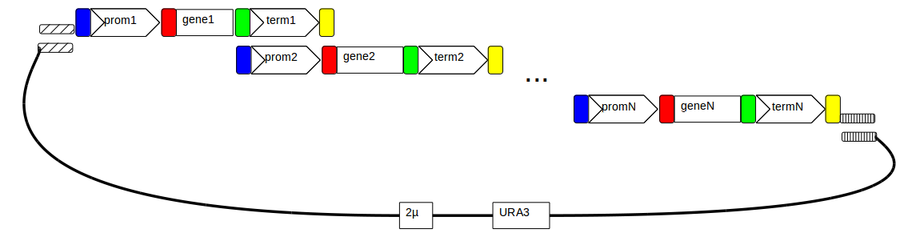
This notebook describes the assembly of 2 single gene expression cassettes into a single pathway. Notebooks describing the single gene expression vectors are linked at the end of this document as are notebooks describing pYPKa promoter, gene and terminator vectors. Specific primers needed are also listed below.
The pydna package is imported in the code cell below. There is a publication describing pydna as well as documentation available online. Pydna is developed on Github.
The assembly performed here is based on content of the INDATA_pth2.txt text file. The assembly log can be viewed here.
from pydna.parsers import parse_primers
from pydna.readers import read
from pydna.amplify import pcr
from pydna.assembly import Assembly
Initiate the standard primers needed to amplify each cassette. The first cassette in the pathway is amplified with standard primers 577 and 778, the last with 775 and 578 and all others with 775 and 778. Standard primers are listed here.
p = { x.id: x for x in parse_primers("standard_primers.txt") }
The backbone vector is linearized with EcoRV.
from Bio.Restriction import EcoRV, NotI, PacI
pYPKpw = read("pYPKpw.gb")
The cassette_products variable holds the list of expression cassette PCR products fragments to be assembled.
cassette_products = []
The expression cassettes comes from a series of single gene expression vectors held in the template_vectors list.
cassette_vectors ='''
pYPK0_TEF1_SsXYL1_TDH3.gb
pYPK0_TDH3_SsXYL2_PGI.gb'''.splitlines()
template_vectors = [read(v.strip()) for v in cassette_vectors if v.strip()]
template_vectors
[File(id?)(o8024), File(id?)(o8580)]
The first cassette in the pathway is amplified with standard primers 577 and 778. Suggested PCR conditions can be found at the end of this document.
cassette_products.append( pcr( p['577'], p['778'], template_vectors[0] ) )
Cassettes in the middle cassettes are amplified with standard primers 775 and 778. Suggested PCR conditions can be found at the end of this document.
cassette_products.extend( pcr( p['775'], p['778'], v) for v in template_vectors[1:-1] )
The last cassette in the pathway is amplified with standard primers 775 and 578. Suggested PCR conditions can be found at the end of this document.
cassette_products.append( pcr( p['775'], p['578'], template_vectors[-1] ) )
The cassettes are given names based on their order in the final construct in the code cell below.
for i, cp in enumerate(cassette_products):
cp.name = "Cassette {}".format(i+1)
print(cp.name)
Cassette 1 Cassette 2
Cassettes and plasmid backbone are joined by homologous recombination in a Saccharomyces cerevisiae ura3 host which selects for the URA3 gene in pYPKpw.
asm = Assembly( [pYPKpw.linearize(EcoRV)] + cassette_products, limit=167-47-10)
asm
Assembly: Sequences........................: [5603] [2524] [3206] Sequences with shared homologies.: [5603] [2524] [3206] Homology limit (bp)..............: 110 Number of overlaps...............: 3 Nodes in graph(incl. 5' & 3')....: 5 Only terminal overlaps...........: No Circular products................: [10212] Linear products..................: [10948] [10456] [10353] [8565] [7986] [4994] [736] [244] [141]
Normally, only one circular product should be formed since the homology limit is quite large (see cell above). More than one circular products might indicate an incorrect strategy. The largest recombination product is chosen as candidate for the pYPK0_SsXYL1_SsXYL2 pathway.
candidate = asm.circular_products[0]
This assembly figure shows how the fragments came together.
candidate.figure()
-|pYPKpw|124 | \/ | /\ | 124|Cassette 1|711 | \/ | /\ | 711|Cassette 2|242 | \/ | /\ | 242- | | ------------------------------------------
The final pathway is synchronized to the backbone vector. This means that the plasmid origin is shifted so that it matches the original.
pw = candidate.synced(pYPKpw)
The cseguid checksum for the resulting plasmid is calculated for future reference. The cseguid checksum uniquely identifies a circular double stranded sequence.
pw.cseguid()
rI2M08mHNu-8plq5MeA8B1K9-mc
The file is given a name based on the sequence of expressed genes.
pw.locus = "pw"
pw.definition = "pYPK0_SsXYL1_SsXYL2"
Stamp sequence with cseguid checksum. This can be used to verify the integrity of the sequence file.
pw.stamp()
cSEGUID_rI2M08mHNu-8plq5MeA8B1K9-mc
Write sequence to a local file.
pw.write("pYPK0_SsXYL1_SsXYL2.gb")
The pathway can be extended by digestion with either NotI or PacI or both provided that the enzymes cut once in the final pathway sequence.
print("NotI cuts {} time(s) and PacI cuts {} time(s) in the final pathway.".format(len(pw.cut(NotI)), len(pw.cut(PacI))))
NotI cuts 1 time(s) and PacI cuts 1 time(s) in the final pathway.
DOWNLOAD pYPK0_SsXYL1_SsXYL2¶
import pydna
reloaded = read("pYPK0_SsXYL1_SsXYL2.gb")
reloaded.verify_stamp()
cSEGUID_rI2M08mHNu-8plq5MeA8B1K9-mc
try:
with open("new_primers.txt") as f:
text = f.read()
except IOError:
text = "no new primers needed."
print(text)
>fw1000 PGI ttaaatAATTCAGTTTTCTGACTGAGT >rv1000 PGI taattaaTTTTAGGCTGGTATCTTGAT >fw1000 PGI ttaaatAATTCAGTTTTCTGACTGAGT >rv1000 PGI taattaaTTTTAGGCTGGTATCTTGAT >fw698 TDH3 ttaaatATAAAAAACACGCTTTTTC >rv698 TDH3 taattaaTTTGTTTGTTTATGTGTGTTT >fw698 TDH3 ttaaatATAAAAAACACGCTTTTTC >rv698 TDH3 taattaaTTTGTTTGTTTATGTGTGTTT >fw579 TEF1 ttaaatACAATGCATACTTTGTACGT >rv579 TEF1 taattaaTTTGTAATTAAAACTTAGATTAGATTG >fw579 TEF1 ttaaatACAATGCATACTTTGTACGT >rv579 TEF1 taattaaTTTGTAATTAAAACTTAGATTAGATTG >fw1092 SsXYL2 tgcccactttctcactagtgacctgcagccgacAAATGACTGCTAACCCTTCC >rv1092 SsXYL2 AAatcctgatgcgtttgtctgcacagatggCACTTACTCAGGGCCGTCA >fw1092 SsXYL2 tgcccactttctcactagtgacctgcagccgacAAATGACTGCTAACCCTTCC >rv1092 SsXYL2 AAatcctgatgcgtttgtctgcacagatggCACTTACTCAGGGCCGTCA >fw957 SsXYL1 tgcccactttctcactagtgacctgcagccgacAAATGCCTTCTATTAAGTTGAAC >rv957 SsXYL1 AAatcctgatgcgtttgtctgcacagatggCACTTAGACGAAGATAGGAATCTTG >fw957 SsXYL1 tgcccactttctcactagtgacctgcagccgacAAATGCCTTCTATTAAGTTGAAC >rv957 SsXYL1 AAatcctgatgcgtttgtctgcacagatggCACTTAGACGAAGATAGGAATCTTG
New single gene expression vectors (pYPK0_prom_gene_term) needed for assembly.¶
Hyperlinks to notebook files describing the singlke gene expression plasmids needed for the assembly.
pYPK0_TEF1_SsXYL1_TDH3
pYPK0_TDH3_SsXYL2_PGI
New pYPKa vectors needed for assembly of the single gene expression vectors above.¶
Hyperlinks to notebook files describing the pYPKa plasmids needed for the assembly of the single gene clones listed above.
pYPKa_ZE_TEF1
pYPKa_ZE_TDH3
pYPKa_ZE_PGI
Suggested PCR conditions¶
for prd in cassette_products:
print("\n\n\n\n")
print("product name:", prd.name)
print("forward primer", prd.forward_primer.name)
print("reverse primer", prd.reverse_primer.name)
print(prd.program())
product name: Cassette 1 forward primer 577 reverse primer 778 Taq (rate 30 nt/s) 35 cycles |2524bp 95.0°C |95.0°C | |Tm formula: Biopython Tm_NN |_________|_____ 72.0°C |72.0°C|SaltC 50mM | 03min00s|30s \ ________|______|Primer1C 1.0µM | | \ 53.1°C/ 1min16s| 5min |Primer2C 1.0µM | | \_____/ | |GC 40% | | 30s | |4-12°C product name: Cassette 2 forward primer 775 reverse primer 578 Taq (rate 30 nt/s) 35 cycles |3206bp 95.0°C |95.0°C | |Tm formula: Biopython Tm_NN |_________|_____ 72.0°C |72.0°C|SaltC 50mM | 03min00s|30s \ ________|______|Primer1C 1.0µM | | \ 56.9°C/ 1min36s| 5min |Primer2C 1.0µM | | \_____/ | |GC 42% | | 30s | |4-12°C