pYPK0_FBA1_ScTAL1_PDC1¶
This notebook describes the assembly of the Saccaromyces cerevisiae single gene expression vector pYPK0_FBA1_ScTAL1_PDC1.
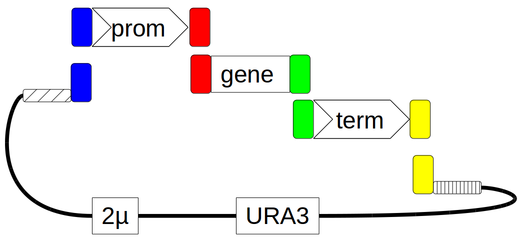
It is made by in-vivo homologous recombination between three PCR products and one linear vector fragment. The PCR products are a promoter generated from a pYPK_Z vector, a gene from a template and a terminator from a pYPKa_E vector. The three PCR products are joined to a linearized pYPKpw backbone vector that has the URA3 marker and a S. crevisiae 2 micron origin of replication.
The four linear DNA fragments are joined by homologous recombination in a Saccharomyces cerevisiae ura3 mutant.
The pydna package is imported in the code cell below. There is a publication describing pydna as well as documentation available online. Pydna is developed on Github.
from pydna.parsers import parse_primers
from pydna.readers import read
from pydna.design import primer_design
from pydna.amplify import pcr
from pydna.assembly import Assembly
The YPK standard primers are read into a dictionary in the code cell below.
p = { x.id: x for x in parse_primers("standard_primers.txt") }
The backbone vector pYPKpw is read from a local file in the code cell below.
pYPKpw = read("pYPKpw.gb")
from Bio.Restriction import EcoRV
pYPK_EcoRV = pYPKpw.linearize(EcoRV)
The pYPKa derived E. coli plasmids containing the promoter and terminator as well as the gene template sequence are read into three variables in the code cell below.
promoter_template = read("pYPKa_Z_FBA1.gb")
gene_template = read("ScTAL1.gb")
terminator_template = read("pYPKa_E_PDC1.gb")
The construction of the two vector above are described in the pYPKa_ZE_FBA1 notebooks.
The promoter is amplified with from pYPKa_Z_FBA1. A suggested PCR program can be found at the end of this document.
prom = pcr( p['577'], p['567'], promoter_template)
Primers with tails are needed for the recombination between the gene and the promoter/terminator PCR products. The tails are designed to provide 33 bp of terminal homology between the DNA fragments.
fp_tail = "tgcccactttctcactagtgacctgcagccgacAA"
rp_tail = "AAatcctgatgcgtttgtctgcacagatggCAC"
Primers with the tails above are designed for the gene template in the code cell below.
ins = primer_design(gene_template)
fp = fp_tail + ins.forward_primer
rp = rp_tail + ins.reverse_primer
The primers are included in the new_primers.txt list and in the end of the pathway notebook file.
print(fp.format("fasta"))
print(rp.format("fasta"))
with open("new_primers.txt", "a+") as f:
f.write(fp.format("fasta"))
f.write(rp.format("fasta"))
>fw1008 ScTAL1 tgcccactttctcactagtgacctgcagccgacAAATGTCTGAACCAGCTCA >rv1008 ScTAL1 AAatcctgatgcgtttgtctgcacagatggCACTTAAGCGGTAACTTTCTTTTC
The gene is amplifed using the newly designed primers. A suggested PCR program can be found at the end of this document.
gene = pcr( fp, rp, gene_template)
The terminator is amplified from pYPKa_E_PDC1. A suggested PCR program can be found at the end of this document.
term = pcr( p['568'], p['578'], terminator_template)
The four linear DNA fragments are mixed and transformed to a Saccharomyces cerevisiae ura3 mutant.
The fragments will be assembled by in-vivo homologous recombination:
asm = Assembly( (pYPK_EcoRV, prom, gene, term), limit=31 )
asm
Assembly: Sequences........................: [5603] [861] [1076] [1294] Sequences with shared homologies.: [5603] [861] [1294] [1076] Homology limit (bp)..............: 31 Number of overlaps...............: 4 Nodes in graph(incl. 5' & 3')....: 6 Only terminal overlaps...........: No Circular products................: [8383] Linear products..................: [8627] [8524] [8416] [8416] [7696] [7373] [7366] [6653] [6323] [3165] [2337] [1904] [244] [141] [33] [33]
The representation of the asm object above should normally indicate one circcular product only.
More than one circular products might indicate an incorrect assembly strategy or represent
by-products that might arise in the assembly process.
The largest recombination product is chosen as candidate for the pYPK0_FBA1_ScTAL1_PDC1 vector.
candidate = asm.assemble_circular()[0]
candidate.figure()
-|pYPKpw|124 | \/ | /\ | 124|861bp_PCR_prod|33 | \/ | /\ | 33|1076bp_PCR_prod|33 | \/ | /\ | 33|1294bp_PCR_prod|242 | \/ | /\ | 242- | | ---------------------------------------------------------------------
The candidate vector is synchronized to the backbone vector. This means that the plasmid origin is shifted so that it matches the pYPKpw backbone vector.
result = candidate.synced(pYPKpw)
Diagnostic PCR confirmation¶
The structure of the final vector is confirmed by two separate PCR reactions, one for the promoter and gene and one for the gene and terminator.
PCR using standard primers 577 and 467 to amplify promoter and gene.
product = pcr( p['577'], p['467'], result)
A correct clone should give this size in base pairs:
print(len(product))
1908
If the promoter is missing from the assembly, the PCR product will have this size in base pairs:
print(len(product) - len(prom))
1047
If the gene is missing from the assembly, the PCR product will have this size in base pairs:
print(len(product) - len(gene))
832
PCR using standard primers 468 and 578 to amplify gene and terminator.
product2 = pcr( p['468'], p['578'], result)
A correct clone should give this size:
print(len(product2))
2354
If the gene is missing from the assembly, the PCR product will have this size in base pairs:
print(len(product2) - len(gene))
1278
If the terminator is missing from the assembly, the PCR product will have this size in base pairs:
print(len(product2) - len(term))
1060
The cseguid checksum for the resulting plasmid is calculated for future reference. The cseguid checksum uniquely identifies a circular double stranded sequence.
result.cseguid()
sdxw3soX7OnU1J90NW60ANRVFdM
The file is named based on the names of promoter, gene and terminator.
result.locus = "pYPK0_tp_g_tp"
result.definition = "pYPK0_FBA1_ScTAL1_PDC1"
Sequence is stamped with cseguid checksum. This can be used to verify the integrity of the sequence file.
result.stamp()
cSEGUID_sdxw3soX7OnU1J90NW60ANRVFdM
Write sequence to a local file.
result.write("pYPK0_FBA1_ScTAL1_PDC1.gb")
PCR programs for the amplification of Promoter, Gene and Terminator¶
Promoter
prom.program()
Taq (rate 30 nt/s) 35 cycles |861bp 95.0°C |95.0°C | |Tm formula: Biopython Tm_NN |_________|_____ 72.0°C |72.0°C|SaltC 50mM | 03min00s|30s \ ________|______|Primer1C 1.0µM | | \ 55.9°C/ 0min26s| 5min |Primer2C 1.0µM | | \_____/ | |GC 39% | | 30s | |4-12°C
Gene
gene.program()
Taq (rate 30 nt/s) 35 cycles |1076bp 95.0°C |95.0°C | |Tm formula: Biopython Tm_NN |_________|_____ 72.0°C |72.0°C|SaltC 50mM | 03min00s|30s \ ________|______|Primer1C 1.0µM | | \ 54.5°C/ 0min32s| 5min |Primer2C 1.0µM | | \_____/ | |GC 42% | | 30s | |4-12°C
Terminator
term.program()
Taq (rate 30 nt/s) 35 cycles |1294bp 95.0°C |95.0°C | |Tm formula: Biopython Tm_NN |_________|_____ 72.0°C |72.0°C|SaltC 50mM | 03min00s|30s \ ________|______|Primer1C 1.0µM | | \ 55.9°C/ 0min39s| 5min |Primer2C 1.0µM | | \_____/ | |GC 38% | | 30s | |4-12°C
Download pYPK0_FBA1_ScTAL1_PDC1¶
import pydna
reloaded = read("pYPK0_FBA1_ScTAL1_PDC1.gb")
reloaded.verify_stamp()
cSEGUID_sdxw3soX7OnU1J90NW60ANRVFdM