2 COUPLED reactions of different speeds, forming a "cascade":¶
A <-> B (fast) and B <-> C (slow)¶
Taken to equilibrium. Both reactions are mostly forward. All 1st order.
The concentration of the intermediate product B manifests 1 oscillation (transient "overshoot")
Adaptive variable time resolution is used, with extensive diagnostics, and finally compared to a new run using fixed time intervals, with the same initial data.
In part2, some diagnostic insight is explored.
In part3, two identical runs ("adaptive variable steps" vs. "fixed small steps") are compared.
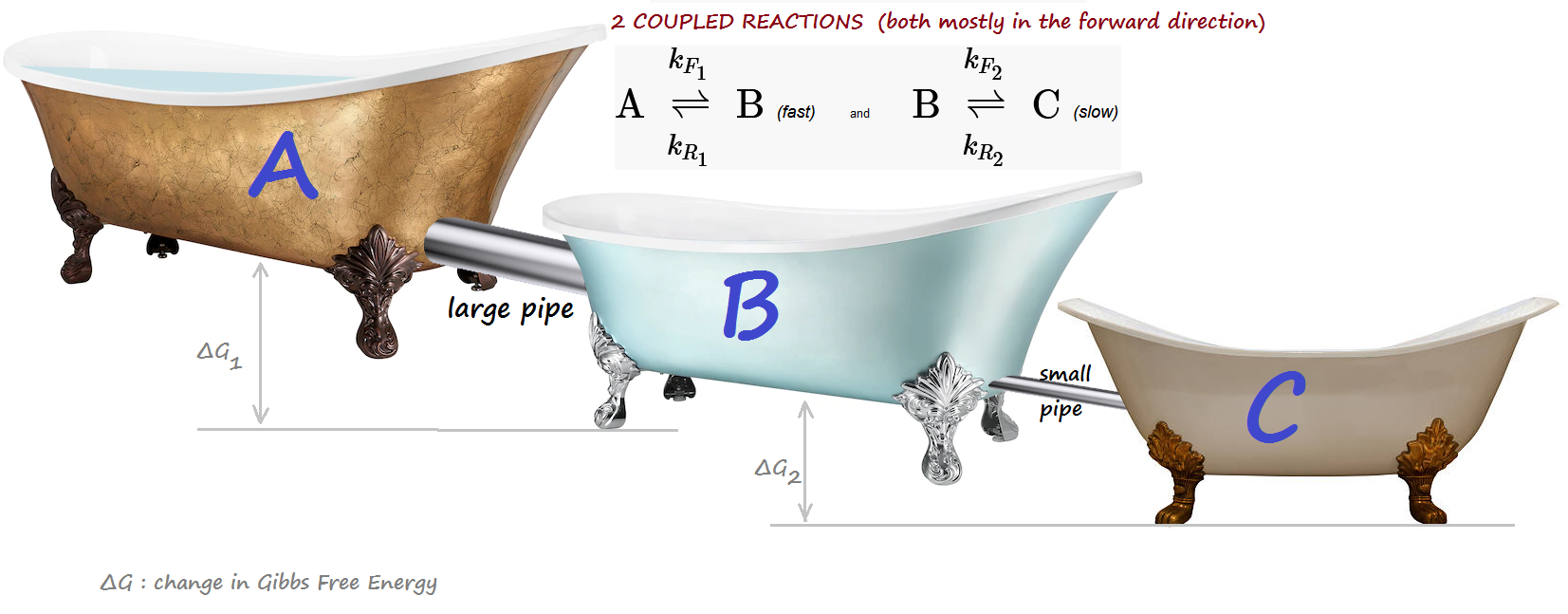
Bathtub analogy:¶
A is initially full, while B and C are empty.
Tubs are progressively lower (reactions are mostly forward.)
A BIG pipe connects A and B: fast kinetics. A small pipe connects B and C: slow kinetics.
INTUITION: B, unable to quickly drain into C while at the same time being blasted by a hefty inflow from A,
will experience a transient surge, in excess of its final equilibrium level.
LAST_REVISED = "Oct. 11, 2024"
LIFE123_VERSION = "1.0.0.beta.39" # Library version this experiment is based on
#import set_path # Using MyBinder? Uncomment this before running the next cell!
#import sys
#sys.path.append("C:/some_path/my_env_or_install") # CHANGE to the folder containing your venv or libraries installation!
# NOTE: If any of the imports below can't find a module, uncomment the lines above, or try: import set_path
import ipynbname
from life123 import check_version, ChemData, UniformCompartment, PlotlyHelper, GraphicLog
check_version(LIFE123_VERSION)
OK
# Initialize the HTML logging (for the graphics)
log_file = ipynbname.name() + ".log.htm" # Use the notebook base filename for the log file
# Set up the use of some specified graphic (Vue) components
GraphicLog.config(filename=log_file,
components=["vue_cytoscape_2"],
extra_js="https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js")
-> Output will be LOGGED into the file 'cascade_1.log.htm'
Initialize the System¶
Specify the chemicals and the reactions
# Specify the chemicals and the reactions; this data structure will get re-used in
# the various simulations below
chem = ChemData()
# Reaction A <-> B (fast)
chem.add_reaction(reactants="A", products="B",
forward_rate=64., reverse_rate=8.)
# Reaction B <-> C (slow)
chem.add_reaction(reactants="B", products="C",
forward_rate=12., reverse_rate=2.)
print("Number of reactions: ", chem.number_of_reactions())
Number of reactions: 2
chem.describe_reactions()
Number of reactions: 2 (at temp. 25 C)
0: A <-> B (kF = 64 / kR = 8 / delta_G = -5,154.8 / K = 8) | 1st order in all reactants & products
1: B <-> C (kF = 12 / kR = 2 / delta_G = -4,441.7 / K = 6) | 1st order in all reactants & products
Set of chemicals involved in the above reactions: {'C', 'A', 'B'}
# Send a plot of the network of reactions to the HTML log file
chem.plot_reaction_network("vue_cytoscape_2")
[GRAPHIC ELEMENT SENT TO LOG FILE `cascade_1.log.htm`]
Run the simulation¶
dynamics = UniformCompartment(chem_data=chem, preset="fast")
dynamics.set_conc({"A": 50.}, snapshot=True) # Set the initial concentrations of all the chemicals, in their index order
dynamics.describe_state()
SYSTEM STATE at Time t = 0:
3 species:
Species 0 (A). Conc: 50.0
Species 1 (B). Conc: 0.0
Species 2 (C). Conc: 0.0
Set of chemicals involved in reactions: {'C', 'A', 'B'}
dynamics.get_history()
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.0 | 50.0 | 0.0 | 0.0 | Initialized state |
Run the reaction¶
dynamics.enable_diagnostics() # To save diagnostic information about the call to single_compartment_react()
dynamics.single_compartment_react(initial_step=0.02, duration=0.4,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
variable_steps=True)
Some steps were backtracked and re-done, to prevent negative concentrations or excessively large concentration changes
48 total step(s) taken
Number of step re-do's because of negative concentrations: 1
Number of step re-do's because of elective soft aborts: 5
Norm usage: {'norm_A': 25, 'norm_B': 17, 'norm_C': 16, 'norm_D': 16}
Plots of changes of concentration with time¶
Notice the variable time steps (vertical dashed lines)
dynamics.plot_history(title="Coupled reactions A <-> B and B <-> C",
colors=['darkturquoise', 'orange', 'green'], show_intervals=True)
dynamics.curve_intersect("A", "B", t_start=0, t_end=0.05)
(0.011857732428358053, 24.047031609298816)
dynamics.curve_intersect("A", "C", t_start=0, t_end=0.05)
(0.031934885008449314, 8.852813807230596)
dynamics.curve_intersect("B", "C", t_start=0.05, t_end=0.1)
(0.07740303497735611, 23.285274847029168)
dynamics.get_history()
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.000000 | 50.000000 | 0.000000 | 0.000000 | Initialized state |
| 1 | 0.000778 | 47.511680 | 2.488320 | 0.000000 | 1st reaction step |
| 2 | 0.001400 | 45.632475 | 4.348950 | 0.018575 | |
| 3 | 0.002022 | 43.837347 | 6.111636 | 0.051017 | |
| 4 | 0.002519 | 42.465438 | 7.447097 | 0.087465 | |
| 5 | 0.003017 | 41.142542 | 8.725606 | 0.131851 | |
| 6 | 0.003515 | 39.866871 | 9.949300 | 0.183829 | |
| 7 | 0.004012 | 38.636703 | 11.120234 | 0.243063 | |
| 8 | 0.004510 | 37.450378 | 12.240391 | 0.309231 | |
| 9 | 0.005008 | 36.306298 | 13.311680 | 0.382022 | |
| 10 | 0.005505 | 35.202922 | 14.335939 | 0.461139 | |
| 11 | 0.006003 | 34.138767 | 15.314939 | 0.546294 | |
| 12 | 0.006501 | 33.112404 | 16.250386 | 0.637210 | |
| 13 | 0.006998 | 32.122455 | 17.143922 | 0.733623 | |
| 14 | 0.007496 | 31.167595 | 17.997130 | 0.835276 | |
| 15 | 0.008243 | 29.786018 | 19.218736 | 0.995246 | |
| 16 | 0.008989 | 28.477742 | 20.356337 | 1.165921 | |
| 17 | 0.009736 | 27.238764 | 21.414705 | 1.346531 | |
| 18 | 0.010482 | 26.065300 | 22.398347 | 1.536353 | |
| 19 | 0.011602 | 24.398010 | 23.768113 | 1.833877 | |
| 20 | 0.012722 | 22.862474 | 24.988386 | 2.149140 | |
| 21 | 0.013841 | 21.447911 | 26.071994 | 2.480095 | |
| 22 | 0.014961 | 20.144428 | 27.030704 | 2.824868 | |
| 23 | 0.016641 | 18.342204 | 28.297603 | 3.360193 | |
| 24 | 0.018320 | 16.750734 | 29.330012 | 3.919255 | |
| 25 | 0.020000 | 15.344212 | 30.158542 | 4.497247 | |
| 26 | 0.022519 | 13.477920 | 31.135708 | 5.386372 | |
| 27 | 0.025039 | 11.932250 | 31.767190 | 6.300559 | |
| 28 | 0.027558 | 10.648537 | 32.122231 | 7.229232 | |
| 29 | 0.031337 | 9.044186 | 32.324491 | 8.631323 | |
| 30 | 0.035116 | 7.833986 | 32.134026 | 10.031989 | |
| 31 | 0.038896 | 6.910732 | 31.675838 | 11.413430 | |
| 32 | 0.044564 | 5.840026 | 30.721211 | 13.438763 | |
| 33 | 0.050233 | 5.114478 | 29.509327 | 15.376196 | |
| 34 | 0.058736 | 4.338557 | 27.535703 | 18.125740 | |
| 35 | 0.065538 | 3.948219 | 25.924919 | 20.126862 | |
| 36 | 0.075742 | 3.486129 | 23.623393 | 22.890478 | |
| 37 | 0.083905 | 3.207568 | 21.961627 | 24.830805 | |
| 38 | 0.096149 | 2.845240 | 19.705148 | 27.449612 | |
| 39 | 0.105945 | 2.605699 | 18.166190 | 29.228112 | |
| 40 | 0.120638 | 2.290745 | 16.137006 | 31.572249 | |
| 41 | 0.135332 | 2.033442 | 14.476842 | 33.489716 | |
| 42 | 0.157371 | 1.717708 | 12.439976 | 35.842316 | |
| 43 | 0.179411 | 1.488195 | 10.959299 | 37.552505 | |
| 44 | 0.212471 | 1.237929 | 9.344771 | 39.417300 | |
| 45 | 0.245531 | 1.090175 | 8.391543 | 40.518282 | |
| 46 | 0.295121 | 0.959315 | 7.547371 | 41.493315 | |
| 47 | 0.369506 | 0.883653 | 7.059061 | 42.057286 | |
| 48 | 0.481083 | 0.874584 | 7.001835 | 42.123581 | last reaction step |
dynamics.diagnostics.explain_time_advance()
From time 0 to 0.0007776, in 1 step of 0.000778 From time 0.0007776 to 0.002022, in 2 steps of 0.000622 From time 0.002022 to 0.007496, in 11 steps of 0.000498 From time 0.007496 to 0.01048, in 4 steps of 0.000746 From time 0.01048 to 0.01496, in 4 steps of 0.00112 From time 0.01496 to 0.02, in 3 steps of 0.00168 From time 0.02 to 0.02756, in 3 steps of 0.00252 From time 0.02756 to 0.0389, in 3 steps of 0.00378 From time 0.0389 to 0.05023, in 2 steps of 0.00567 From time 0.05023 to 0.05874, in 1 step of 0.0085 From time 0.05874 to 0.06554, in 1 step of 0.0068 From time 0.06554 to 0.07574, in 1 step of 0.0102 From time 0.07574 to 0.08391, in 1 step of 0.00816 From time 0.08391 to 0.09615, in 1 step of 0.0122 From time 0.09615 to 0.1059, in 1 step of 0.0098 From time 0.1059 to 0.1353, in 2 steps of 0.0147 From time 0.1353 to 0.1794, in 2 steps of 0.022 From time 0.1794 to 0.2455, in 2 steps of 0.0331 From time 0.2455 to 0.2951, in 1 step of 0.0496 From time 0.2951 to 0.3695, in 1 step of 0.0744 From time 0.3695 to 0.4811, in 1 step of 0.112 (48 steps total)
Check the final equilibrium¶
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium()
0: A <-> B
Final concentrations: [A] = 0.8746 ; [B] = 7.002
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 8.0059
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 8
Discrepancy between the two values: 0.07378 %
Reaction IS in equilibrium (within 1% tolerance)
1: B <-> C
Final concentrations: [B] = 7.002 ; [C] = 42.12
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 6.01608
Formula used: [C] / [B]
2. Ratio of forward/reverse reaction rates: 6
Discrepancy between the two values: 0.268 %
Reaction IS in equilibrium (within 1% tolerance)
True
Let's look at the final concentrations of A and C (i.e., the reactant and product of the composite reaction)¶
A_final = dynamics.get_chem_conc("A")
A_final
0.8745841244150234
C_final = dynamics.get_chem_conc("C")
C_final
42.123581082850514
Their ratio:¶
C_final / A_final
48.16412727709344
As expected the equilibrium constant for the overall reaction A <-> C (approx. 48) is indeed the product of the equilibrium constants of the two elementary reactions (K = 8 and K = 6, respectively) that we saw earlier.¶
PART 2 - DIAGNOSTIC INSIGHT¶
Perform some verification
Take a peek at the diagnostic data saved during the earlier reaction simulation¶
# Concentration increments due to reaction 0 (A <-> B)
# Note that [C] is not affected
dynamics.diagnostics.get_rxn_data(rxn_index=0)
Reaction: A <-> B
| START_TIME | time_step | aborted | Delta A | Delta B | caption | rate | |
|---|---|---|---|---|---|---|---|
| 0 | 0.000000 | 0.020000 | True | NaN | NaN | aborted: neg. conc. in `A` | NaN |
| 1 | 0.000000 | 0.010000 | True | -32.000000 | 32.000000 | aborted: excessive norm value(s) | 3200.000000 |
| 2 | 0.000000 | 0.006000 | True | -19.200000 | 19.200000 | aborted: excessive norm value(s) | 3200.000000 |
| 3 | 0.000000 | 0.003600 | True | -11.520000 | 11.520000 | aborted: excessive norm value(s) | 3200.000000 |
| 4 | 0.000000 | 0.002160 | True | -6.912000 | 6.912000 | aborted: excessive norm value(s) | 3200.000000 |
| 5 | 0.000000 | 0.001296 | True | -4.147200 | 4.147200 | aborted: excessive norm value(s) | 3200.000000 |
| 6 | 0.000000 | 0.000778 | False | -2.488320 | 2.488320 | 3200.000000 | |
| 7 | 0.000778 | 0.000622 | False | -1.879205 | 1.879205 | 3020.840960 | |
| 8 | 0.001400 | 0.000622 | False | -1.795128 | 1.795128 | 2885.686820 | |
| 9 | 0.002022 | 0.000498 | False | -1.371909 | 1.371909 | 2756.697133 | |
| 10 | 0.002519 | 0.000498 | False | -1.322896 | 1.322896 | 2658.211272 | |
| 11 | 0.003017 | 0.000498 | False | -1.275671 | 1.275671 | 2563.317851 | |
| 12 | 0.003515 | 0.000498 | False | -1.230168 | 1.230168 | 2471.885360 | |
| 13 | 0.004012 | 0.000498 | False | -1.186325 | 1.186325 | 2383.787110 | |
| 14 | 0.004510 | 0.000498 | False | -1.144080 | 1.144080 | 2298.901050 | |
| 15 | 0.005008 | 0.000498 | False | -1.103376 | 1.103376 | 2217.109600 | |
| 16 | 0.005505 | 0.000498 | False | -1.064155 | 1.064155 | 2138.299488 | |
| 17 | 0.006003 | 0.000498 | False | -1.026363 | 1.026363 | 2062.361590 | |
| 18 | 0.006501 | 0.000498 | False | -0.989949 | 0.989949 | 1989.190778 | |
| 19 | 0.006998 | 0.000498 | False | -0.954861 | 0.954861 | 1918.685776 | |
| 20 | 0.007496 | 0.000746 | False | -1.381577 | 1.381577 | 1850.749018 | |
| 21 | 0.008243 | 0.000746 | False | -1.308275 | 1.308275 | 1752.555256 | |
| 22 | 0.008989 | 0.000746 | False | -1.238978 | 1.238978 | 1659.724818 | |
| 23 | 0.009736 | 0.000746 | False | -1.173464 | 1.173464 | 1571.963289 | |
| 24 | 0.010482 | 0.001120 | False | -1.667290 | 1.667290 | 1488.992431 | |
| 25 | 0.011602 | 0.001120 | False | -1.535536 | 1.535536 | 1371.327721 | |
| 26 | 0.012722 | 0.001120 | False | -1.414563 | 1.414563 | 1263.291238 | |
| 27 | 0.013841 | 0.001120 | False | -1.303483 | 1.303483 | 1164.090351 | |
| 28 | 0.014961 | 0.001680 | False | -1.802224 | 1.802224 | 1072.997749 | |
| 29 | 0.016641 | 0.001680 | False | -1.591470 | 1.591470 | 947.520208 | |
| 30 | 0.018320 | 0.001680 | False | -1.406522 | 1.406522 | 837.406853 | |
| 31 | 0.020000 | 0.002519 | False | -1.866292 | 1.866292 | 740.761211 | |
| 32 | 0.022519 | 0.002519 | False | -1.545670 | 1.545670 | 613.501217 | |
| 33 | 0.025039 | 0.002519 | False | -1.283713 | 1.283713 | 509.526499 | |
| 34 | 0.027558 | 0.003779 | False | -1.604351 | 1.604351 | 424.528524 | |
| 35 | 0.031337 | 0.003779 | False | -1.210200 | 1.210200 | 320.231977 | |
| 36 | 0.035116 | 0.003779 | False | -0.923254 | 0.923254 | 244.302890 | |
| 37 | 0.038896 | 0.005669 | False | -1.070706 | 1.070706 | 188.880147 | |
| 38 | 0.044564 | 0.005669 | False | -0.725549 | 0.725549 | 127.992002 | |
| 39 | 0.050233 | 0.008503 | False | -0.775920 | 0.775920 | 91.251951 | |
| 40 | 0.058736 | 0.006802 | False | -0.390338 | 0.390338 | 57.382032 | |
| 41 | 0.065538 | 0.010204 | False | -0.462090 | 0.462090 | 45.286669 | |
| 42 | 0.075742 | 0.008163 | False | -0.278561 | 0.278561 | 34.125104 | |
| 43 | 0.083905 | 0.012244 | False | -0.362328 | 0.362328 | 29.591334 | |
| 44 | 0.096149 | 0.009796 | False | -0.239541 | 0.239541 | 24.454162 | |
| 45 | 0.105945 | 0.014693 | False | -0.314953 | 0.314953 | 21.435190 | |
| 46 | 0.120638 | 0.014693 | False | -0.257304 | 0.257304 | 17.511654 | |
| 47 | 0.135332 | 0.022040 | False | -0.315733 | 0.315733 | 14.325526 | |
| 48 | 0.157371 | 0.022040 | False | -0.229513 | 0.229513 | 10.413518 | |
| 49 | 0.179411 | 0.033060 | False | -0.250266 | 0.250266 | 7.570088 | |
| 50 | 0.212471 | 0.033060 | False | -0.147754 | 0.147754 | 4.469280 | |
| 51 | 0.245531 | 0.049590 | False | -0.130860 | 0.130860 | 2.638855 | |
| 52 | 0.295121 | 0.074385 | False | -0.075662 | 0.075662 | 1.017172 | |
| 53 | 0.369506 | 0.111577 | False | -0.009068 | 0.009068 | 0.081275 |
# Concentration increments due to reaction 1 (B <-> C)
# Also notice that the 0-th row from the A <-> B reaction isn't seen here (start time 0 and step 0.02),
# because that step was aborted early on, BEFORE even getting to THIS reaction
dynamics.diagnostics.get_rxn_data(rxn_index=1)
Reaction: B <-> C
| START_TIME | time_step | aborted | Delta B | Delta C | rate | caption | |
|---|---|---|---|---|---|---|---|
| 0 | 0.000000 | 0.010000 | True | 0.000000 | 0.000000 | 0.000000 | aborted: excessive norm value(s) |
| 1 | 0.000000 | 0.006000 | True | 0.000000 | 0.000000 | 0.000000 | aborted: excessive norm value(s) |
| 2 | 0.000000 | 0.003600 | True | 0.000000 | 0.000000 | 0.000000 | aborted: excessive norm value(s) |
| 3 | 0.000000 | 0.002160 | True | 0.000000 | 0.000000 | 0.000000 | aborted: excessive norm value(s) |
| 4 | 0.000000 | 0.001296 | True | 0.000000 | 0.000000 | 0.000000 | aborted: excessive norm value(s) |
| 5 | 0.000000 | 0.000778 | False | 0.000000 | 0.000000 | 0.000000 | |
| 6 | 0.000778 | 0.000622 | False | -0.018575 | 0.018575 | 29.859840 | |
| 7 | 0.001400 | 0.000622 | False | -0.032442 | 0.032442 | 52.150244 | |
| 8 | 0.002022 | 0.000498 | False | -0.036448 | 0.036448 | 73.237598 | |
| 9 | 0.002519 | 0.000498 | False | -0.044387 | 0.044387 | 89.190237 | |
| 10 | 0.003017 | 0.000498 | False | -0.051978 | 0.051978 | 104.443575 | |
| 11 | 0.003515 | 0.000498 | False | -0.059234 | 0.059234 | 119.023938 | |
| 12 | 0.004012 | 0.000498 | False | -0.066168 | 0.066168 | 132.956683 | |
| 13 | 0.004510 | 0.000498 | False | -0.072791 | 0.072791 | 146.266235 | |
| 14 | 0.005008 | 0.000498 | False | -0.079117 | 0.079117 | 158.976118 | |
| 15 | 0.005505 | 0.000498 | False | -0.085155 | 0.085155 | 171.108992 | |
| 16 | 0.006003 | 0.000498 | False | -0.090917 | 0.090917 | 182.686681 | |
| 17 | 0.006501 | 0.000498 | False | -0.096413 | 0.096413 | 193.730206 | |
| 18 | 0.006998 | 0.000498 | False | -0.101653 | 0.101653 | 204.259814 | |
| 19 | 0.007496 | 0.000746 | False | -0.159970 | 0.159970 | 214.295006 | |
| 20 | 0.008243 | 0.000746 | False | -0.170675 | 0.170675 | 228.634342 | |
| 21 | 0.008989 | 0.000746 | False | -0.180610 | 0.180610 | 241.944203 | |
| 22 | 0.009736 | 0.000746 | False | -0.189822 | 0.189822 | 254.283393 | |
| 23 | 0.010482 | 0.001120 | False | -0.297524 | 0.297524 | 265.707463 | |
| 24 | 0.011602 | 0.001120 | False | -0.315263 | 0.315263 | 281.549606 | |
| 25 | 0.012722 | 0.001120 | False | -0.330954 | 0.330954 | 295.562349 | |
| 26 | 0.013841 | 0.001120 | False | -0.344773 | 0.344773 | 307.903744 | |
| 27 | 0.014961 | 0.001680 | False | -0.535325 | 0.535325 | 318.718715 | |
| 28 | 0.016641 | 0.001680 | False | -0.559062 | 0.559062 | 332.850855 | |
| 29 | 0.018320 | 0.001680 | False | -0.577992 | 0.577992 | 344.121633 | |
| 30 | 0.020000 | 0.002519 | False | -0.889125 | 0.889125 | 352.908006 | |
| 31 | 0.022519 | 0.002519 | False | -0.914188 | 0.914188 | 362.855756 | |
| 32 | 0.025039 | 0.002519 | False | -0.928673 | 0.928673 | 368.605168 | |
| 33 | 0.027558 | 0.003779 | False | -1.402091 | 1.402091 | 371.008309 | |
| 34 | 0.031337 | 0.003779 | False | -1.400666 | 1.400666 | 370.631249 | |
| 35 | 0.035116 | 0.003779 | False | -1.381442 | 1.381442 | 365.544329 | |
| 36 | 0.038896 | 0.005669 | False | -2.025333 | 2.025333 | 357.283191 | |
| 37 | 0.044564 | 0.005669 | False | -1.937433 | 1.937433 | 341.777002 | |
| 38 | 0.050233 | 0.008503 | False | -2.749544 | 2.749544 | 323.359530 | |
| 39 | 0.058736 | 0.006802 | False | -2.001123 | 2.001123 | 294.176957 | |
| 40 | 0.065538 | 0.010204 | False | -2.763615 | 2.763615 | 270.845299 | |
| 41 | 0.075742 | 0.008163 | False | -1.940327 | 1.940327 | 237.699766 | |
| 42 | 0.083905 | 0.012244 | False | -2.618807 | 2.618807 | 213.877913 | |
| 43 | 0.096149 | 0.009796 | False | -1.778500 | 1.778500 | 181.562555 | |
| 44 | 0.105945 | 0.014693 | False | -2.344137 | 2.344137 | 159.538054 | |
| 45 | 0.120638 | 0.014693 | False | -1.917467 | 1.917467 | 130.499569 | |
| 46 | 0.135332 | 0.022040 | False | -2.352600 | 2.352600 | 106.742678 | |
| 47 | 0.157371 | 0.022040 | False | -1.710189 | 1.710189 | 77.595076 | |
| 48 | 0.179411 | 0.033060 | False | -1.864795 | 1.864795 | 56.406583 | |
| 49 | 0.212471 | 0.033060 | False | -1.100982 | 1.100982 | 33.302648 | |
| 50 | 0.245531 | 0.049590 | False | -0.975033 | 0.975033 | 19.661952 | |
| 51 | 0.295121 | 0.074385 | False | -0.563972 | 0.563972 | 7.581818 | |
| 52 | 0.369506 | 0.111577 | False | -0.066295 | 0.066295 | 0.594162 |
PART 3 : Re-run with very small constant steps, and compare with original run¶
We'll use constant steps of size 0.0005 , which is 1/4 of the smallest steps (the "substep" size) previously used in the variable-step run
dynamics2 = UniformCompartment(chem_data=chem) # Re-use the same chemicals and reactions of the previous simulation
dynamics2.set_conc({"A": 50.}, snapshot=True)
# Notice that we're using FIXED steps this time
dynamics2.single_compartment_react(initial_step=0.0005, duration=0.4,
variable_steps=False,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
)
800 total step(s) taken
dynamics2.plot_history(title="Coupled reactions A <-> B and B <-> C , re-run with CONSTANT STEPS",
colors=['darkturquoise', 'orange', 'green'], show_intervals=True)
plot_pandas() NOTICE: Excessive number of vertical lines (801) - only showing 1 every 6 lines
(Notice that the vertical steps are now equally spaced - and that there are so many of them that we're only showing 1 every 6)
dynamics2.curve_intersect(t_start=0, t_end=0.05, chem1="A", chem2="B")
(0.011934802021672202, 24.03028826511689)
dynamics2.curve_intersect(t_start=0, t_end=0.05, chem1="A", chem2="C")
(0.03259520587890533, 9.081194286771336)
dynamics2.curve_intersect(t_start=0.05, t_end=0.1, chem1="B", chem2="C")
(0.07932921476313338, 23.238745930657448)
df2 = dynamics2.get_history()
df2
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.0000 | 50.000000 | 0.000000 | 0.000000 | Initialized state |
| 1 | 0.0005 | 48.400000 | 1.600000 | 0.000000 | 1st reaction step |
| 2 | 0.0010 | 46.857600 | 3.132800 | 0.009600 | |
| 3 | 0.0015 | 45.370688 | 4.600925 | 0.028387 | |
| 4 | 0.0020 | 43.937230 | 6.006806 | 0.055964 | |
| ... | ... | ... | ... | ... | ... |
| 796 | 0.3980 | 0.925494 | 7.329151 | 41.745354 | |
| 797 | 0.3985 | 0.925195 | 7.327221 | 41.747584 | |
| 798 | 0.3990 | 0.924898 | 7.325303 | 41.749800 | |
| 799 | 0.3995 | 0.924602 | 7.323396 | 41.752002 | |
| 800 | 0.4000 | 0.924309 | 7.321501 | 41.754190 | last reaction step |
801 rows × 5 columns
Notice that we now did 800 steps - vs. the 48 of the earlier variable-resolution run!¶
Let's compare some entries with the coarser previous variable-time run¶
Let's compare the plots of [B] from the earlier (variable-step) run, and the latest (high-precision, fixed-step) one:¶
# Earlier run (using variable time steps)
fig1 = dynamics.plot_history(chemicals='B', colors='orange', title="Adaptive variable-step run",
show=True)
# Latest run (high-precision result from fine fixed-resolution run)
fig2 = dynamics2.plot_history(chemicals='B', colors=['violet'], title="Fine fixed-step run",
show=True)
PlotlyHelper.combine_plots(fig_list=[fig1, fig2], title="The 2 runs, contrasted together",
curve_labels=["B (adaptive variable steps)", "B (fixed small steps)"])
They overlap fairly well! The 800 fixed-timestep points vs. the 48 adaptable variable-timestep ones.¶
The adaptive algorithms avoided 752 extra steps of limited benefit...