Introduction to the Insight Toolkit (ITK)¶
Learning Objectives¶
- Learn how to run cells in a Jupyter Notebook
- Run a segmentation example that demonstrates ITK's ability to provide insight into images
- Understand the purpose and capabilities of the toolkit
Jupyter Notebooks¶
These are Jupyter Notebooks, an open-source web application that allows you to create and share documents that contain live code, equations, visualizations and narrative text.
To run cells in the notebook, press shift + enter.
For more information, see the Notebook Help
Insight Into Images¶
In [1]:
import itk
from packaging.version import parse
from importlib.metadata import version
if parse(version('itk')) < parse('5.3'):
raise ValueError("ITK greater than version 5.3.0 is required for this notebook")
In [2]:
from itkwidgets import view
In [3]:
file_name = "data/brainweb165a10f17.mha"
image = itk.imread(file_name, itk.ctype("float"))
view(image, gradient_opacity=0.5)
Out[3]:
<itkwidgets.viewer.Viewer at 0x7fa2755add10>
In [4]:
# Smooth the image
smoothed = itk.curvature_flow_image_filter(
image, number_of_iterations=6, time_step=0.005
)
view(smoothed, gradient_opacity=0.5)
Out[4]:
<itkwidgets.viewer.Viewer at 0x7fa2733fe010>
In [5]:
# Segment the white matter with a 3D region-growing algorithm
confidence_connected = itk.ConfidenceConnectedImageFilter.New(smoothed)
confidence_connected.SetMultiplier(2.5)
confidence_connected.SetNumberOfIterations(5)
confidence_connected.SetInitialNeighborhoodRadius(2)
confidence_connected.SetReplaceValue(255)
confidence_connected.AddSeed([118, 133, 92])
confidence_connected.AddSeed([63, 135, 94])
confidence_connected.AddSeed([63, 157, 90])
confidence_connected.AddSeed([111, 150, 90])
confidence_connected.AddSeed([111, 50, 88])
confidence_connected.Update()
In [7]:
view(
image=smoothed,
label_image=confidence_connected.GetOutput(),
gradient_opacity=0.5
)
Out[7]:
<itkwidgets.viewer.Viewer at 0x7fa2755ad4d0>
History of ITK¶
History¶
In 1999, the US National Institute of Health’s (NIH) National Library of Medicine (NLM) started a project to support the Visible Human Project.

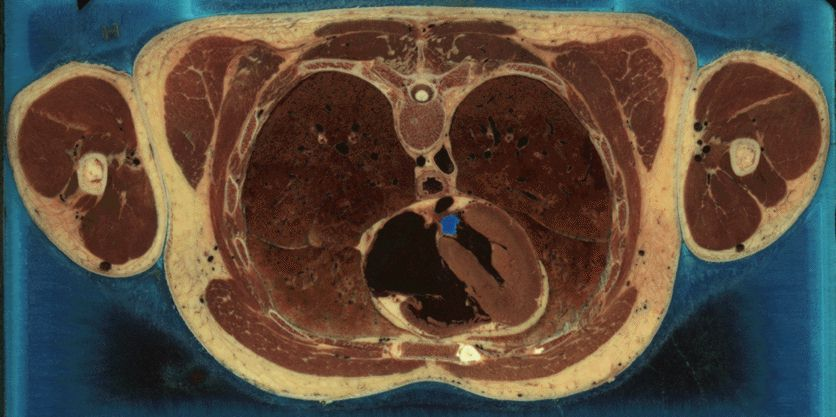
Goals¶
- Collect best-of-the-best image analysis algorithms for reproducible science.
- Provide a software resource for teaching medical image analysis algorithms.
- Establish a foundation for future research.
- Support commercial applications.
- Create conventions for future work.
- Grow a self-sustaining community of software users and developers.
Continued Development¶
- Development has progressed since 1999
- Contributions from over 267 developers
- Over 1.6 million lines of code
- Downloaded over 500,000 times
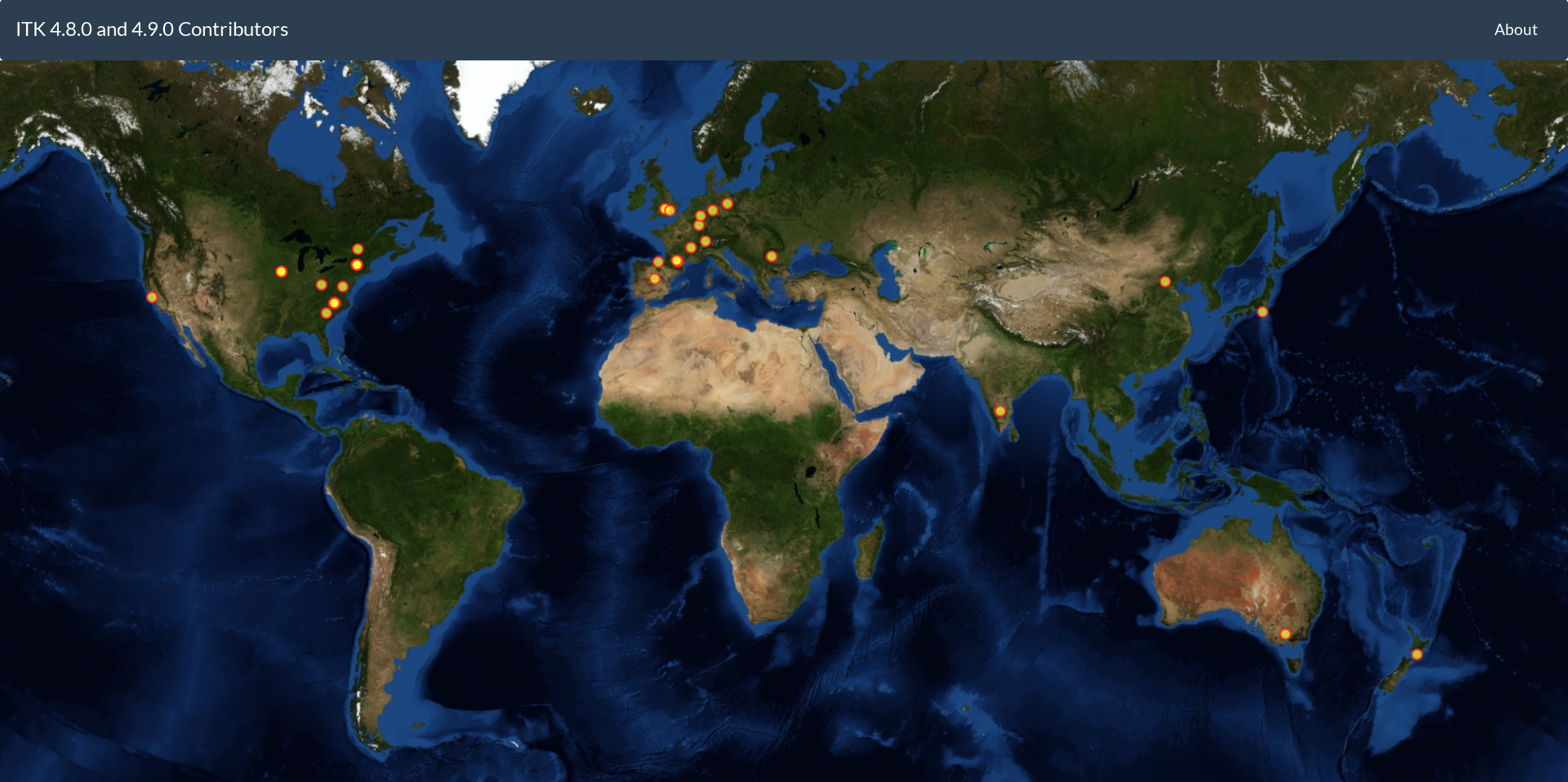
ITK contributors locations for the 4.8 and 4.9 releases.
Features¶
N-Dimensional Image Filtering¶
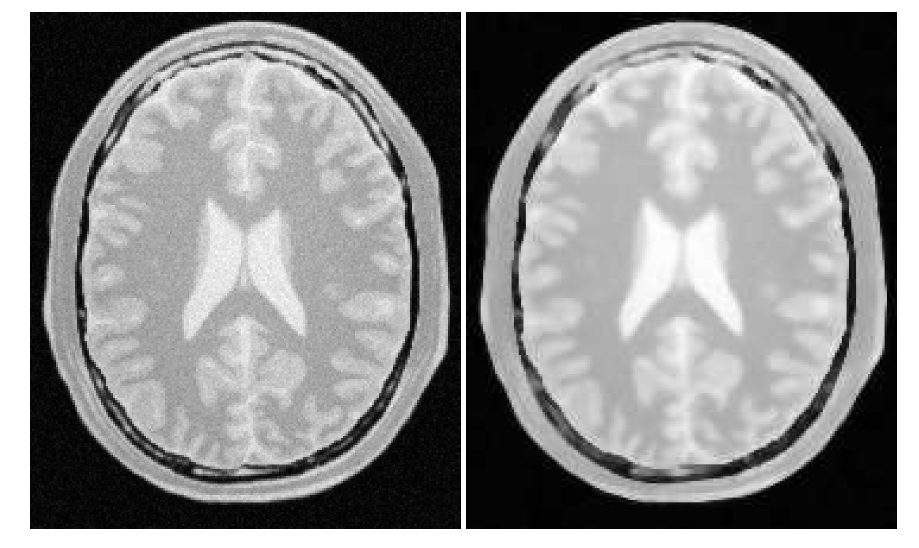
Filtering Algorithms Classes¶
- Fast marching methods
- Convolution
- Image gradient
- Denoising
- Thresholding
- Mathematical morphology
- Smoothing
- Image features
- Image statistics
- Bias correction
- Image grid operations
- ....
Segmentation¶
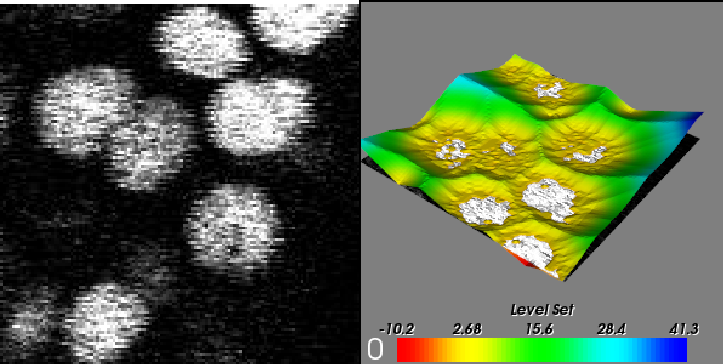
Segmentation¶
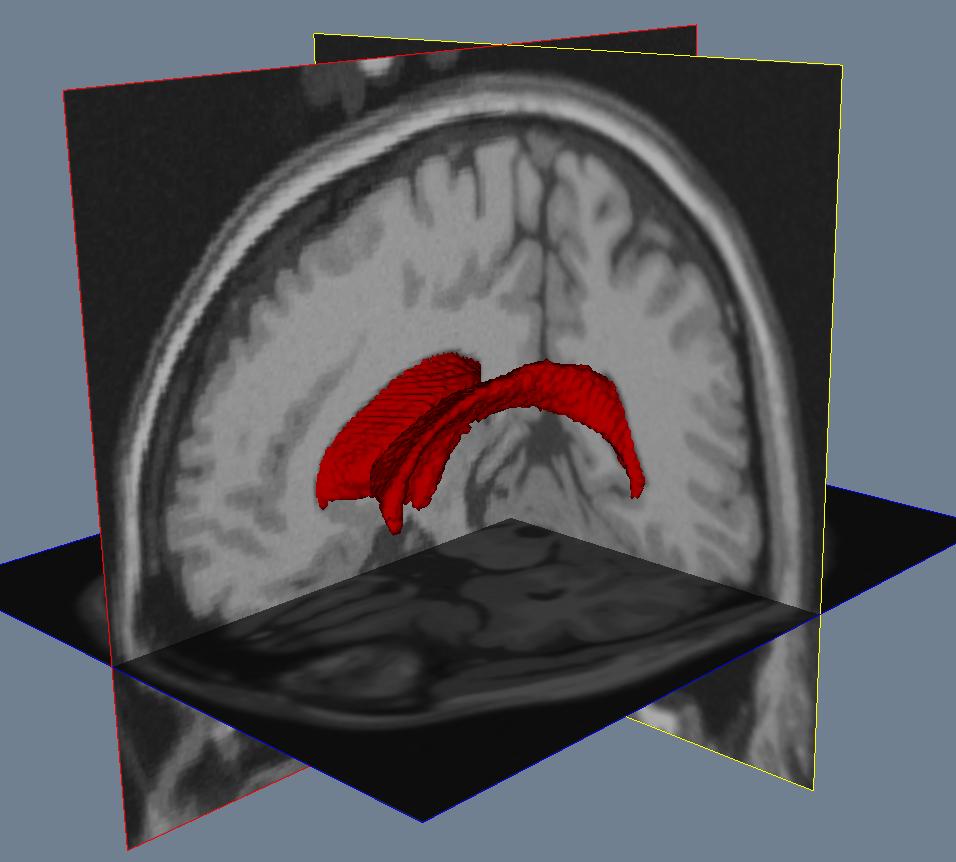
Segmentation¶
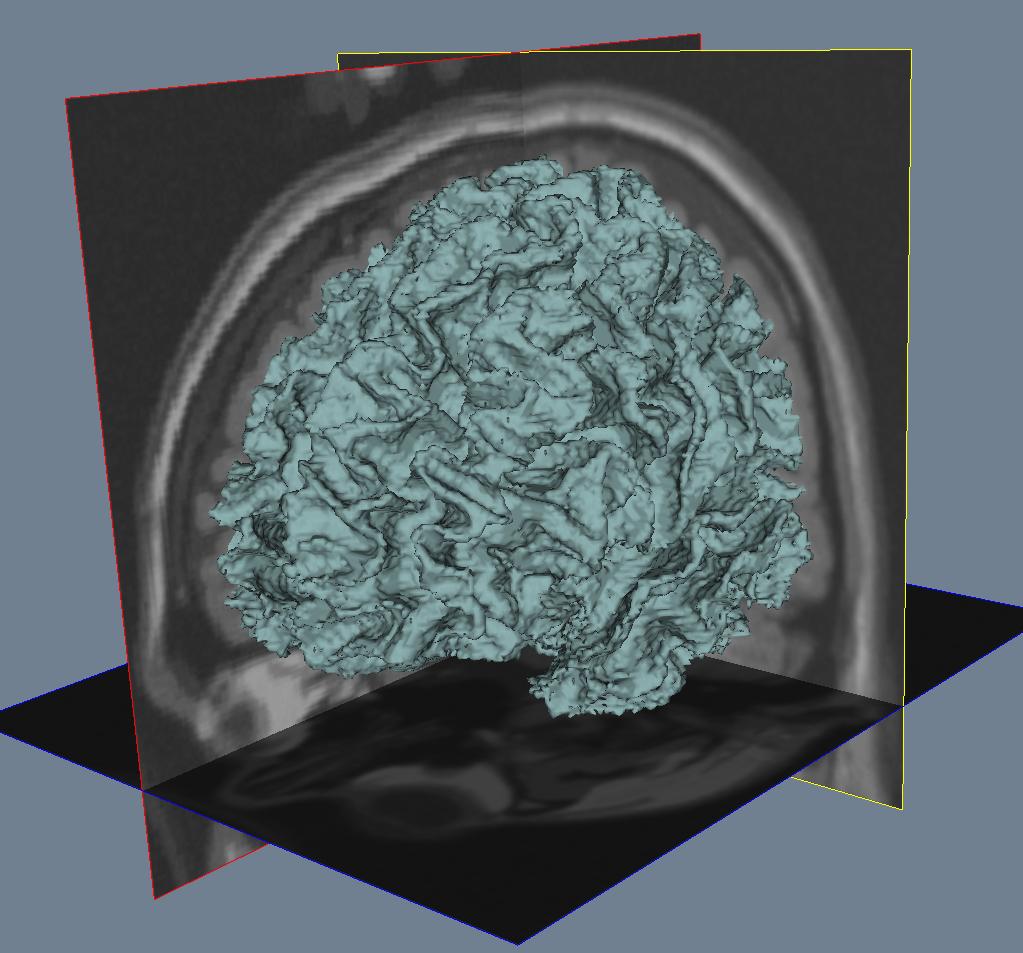
Segmentation¶
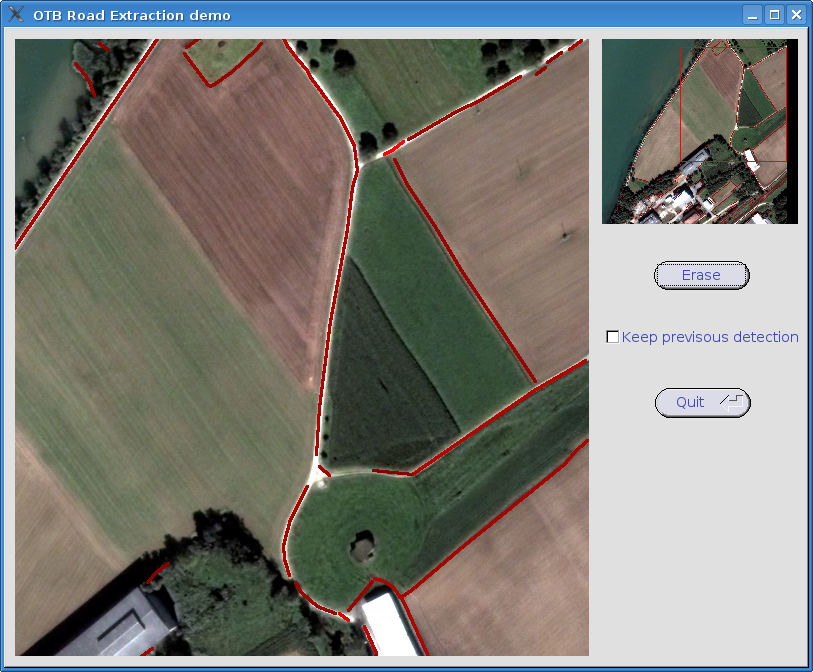
Segmentation¶
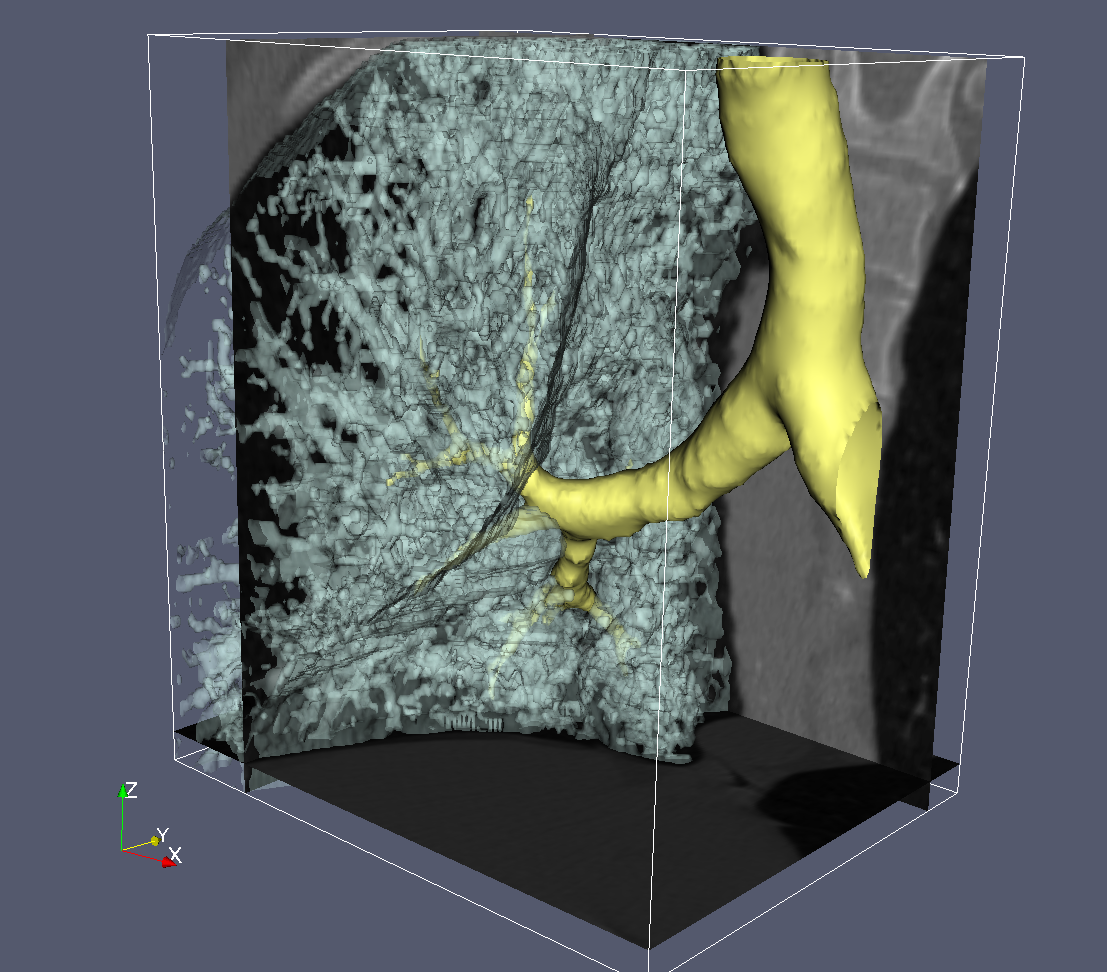

Segmentation¶
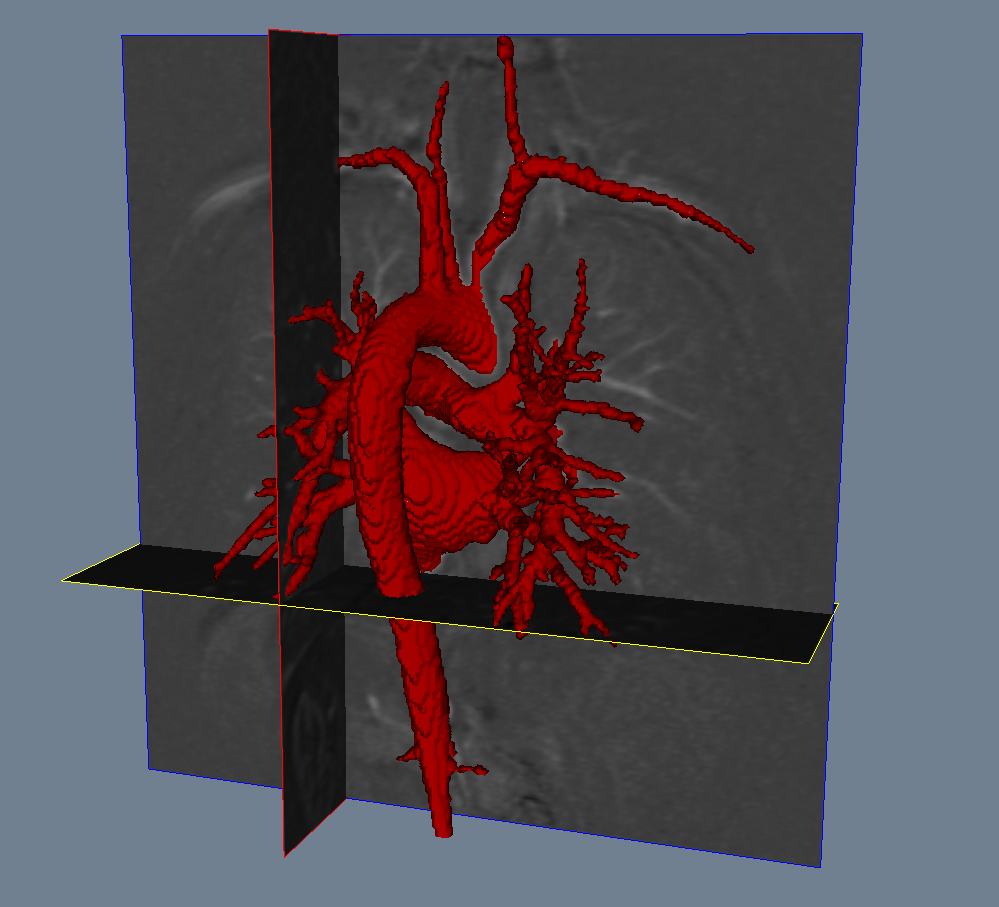
Registration¶
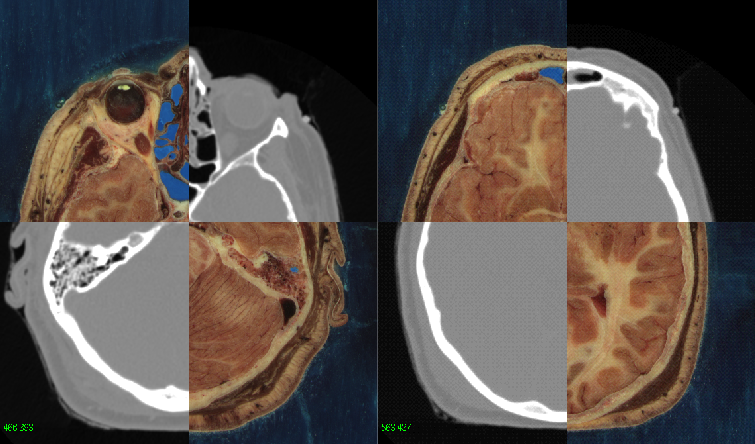
ITK Resources¶
ITK Software Guide: https://www.itk.org/ItkSoftwareGuide.pdf
Discourse Discussion: https://discourse.itk.org
Sphinx Examples: https://www.itk.org/ITKExamples
ITK/Examples/directory in the ITK source code repositoryKitware: https://www.kitware.com/
Kitware, Inc¶


- Collaborative, Open Source, Scientific Software Development
- Carrboro, NC, near University of North Carolina-Chapel Hill, in the Research Triangle