
In [1]:
%load_ext autoreload
%autoreload 2
import sys
sys.path.append("../")
from utils import *
# define paths
input_path = "COVID-19-AR/nifti_files/"
output_path = "COVID-19-AR/output_files/"
cropped_path = "COVID-19-AR/cropped_nifti_files/"
cropped_json_path = "COVID-19-AR/cropped_output_files/"
Preprocess CT scans in COVID-19 dataset¶
1. Download Data¶
Download COVID-19 CT dataset from Zenodo.
The dataset source is the COVID-19-AR data from the Cancer Image Archive.

2. Analyze Data¶
In [2]:
plot_volumes_interactive(input_path, start_index=36)
interactive(children=(BoundedFloatText(value=36.0, description='File:', max=190.0, step=1.0), Output()), _dom_…

3. Define preprocessing steps¶
1. Remove invalid CT scans- filter CT scans, with wrong axis ordering
- filter corrupted CT scans, where no human body is visible
- filter CT scans with less than 20 slices
2. Remove CT scans, where the lungs are not fully visible
3. Crop lungs out of CT scans if more body regions are visible
→ Goal: CT dataset where in all scans the lungs were examined
5. Create body part metadata file for each CT image¶

In [3]:
from bpreg.scripts.bpreg_inference import bpreg_inference
# bpreg_inference(input_path, output_path, plot=True)
6. Analyze body part meta data files¶
In [4]:
plot_scores_interactive(output_path, input_path, start_index=166)
interactive(children=(BoundedFloatText(value=166.0, description='File:', max=190.0, min=166.0, step=1.0), Outp…
In [5]:
df = create_meta_data_table(output_path)
# Filter CT scans with less than 20 slices
df = df[df.z > 20]
plot_dicomexamined_distribution(df,
others_percentage_upper_bound=0.015)
191it [00:00, 470.90it/s]
7. Remove invalid CT scans¶
Filter CT scans, were the predicted body part is NONE
In [6]:
# Filter corrupted CT scans
df_cleaned = df[df['BODY PART'] != 'NONE']
print(f"Dataset size after removing {len(df) - len(df_cleaned)} corrupted CT scans: {len(df_cleaned)}")
Dataset size after removing 20 corrupted CT scans: 125
In [7]:
plot_dicomexamined_distribution(df_cleaned,
others_percentage_upper_bound=0.02)
8. Filter chest CT scans¶
In [8]:
# Removing CT scans where the chest is not examined
df_cleaned = df_cleaned[df_cleaned['BODY PART'].str.contains('CHEST')]
print(f"Dataset after removing CT scans, where the CHEST is not visible: {len(df_cleaned)}")
Dataset after removing CT scans, where the CHEST is not visible: 107
In [9]:
plot_dicomexamined_distribution(df_cleaned,
others_percentage_upper_bound=0.02)
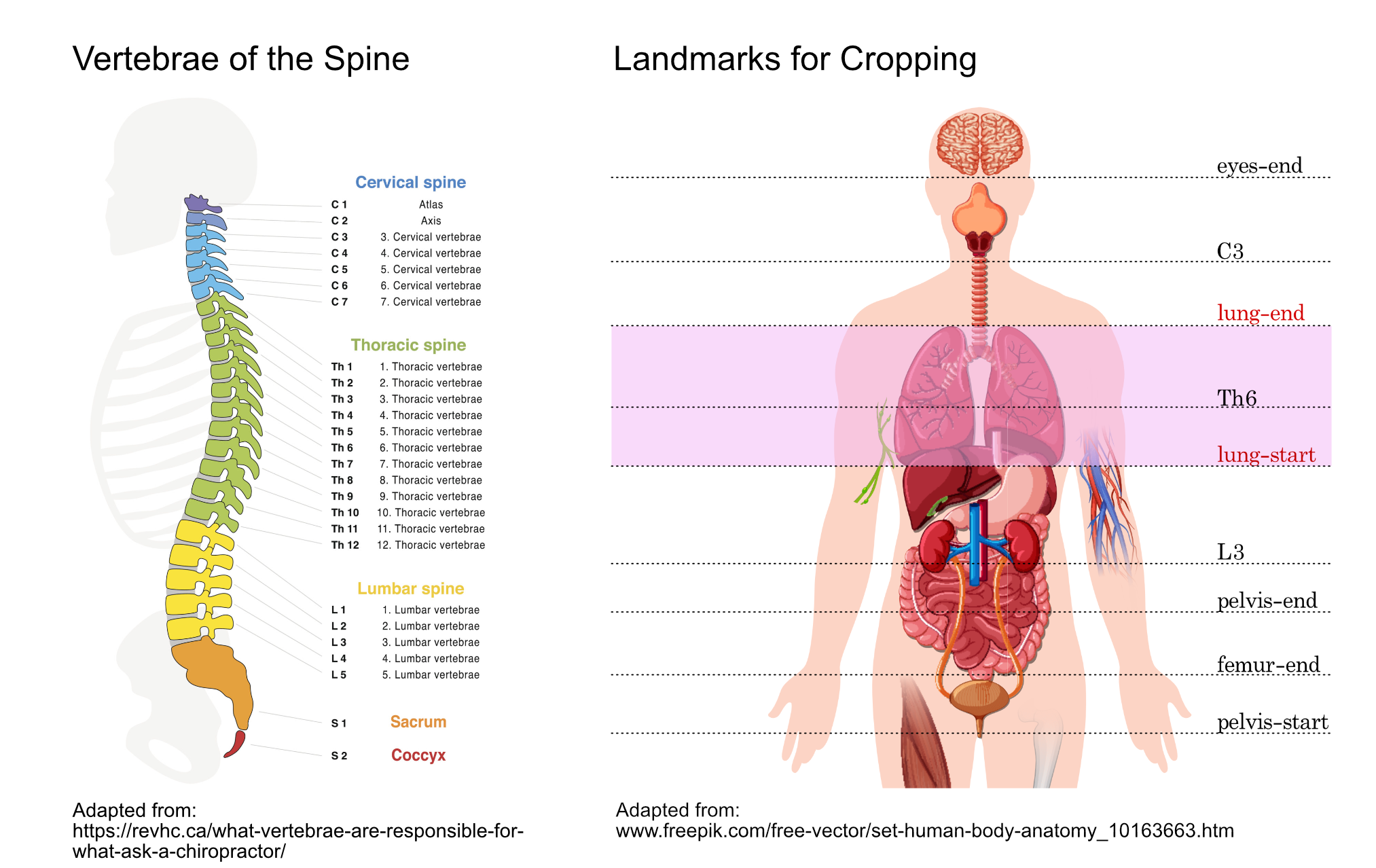
9. Crop chest region out of CT scan¶
In [10]:
import pandas as pd
json_filepaths = [output_path + f for f in os.listdir(output_path) if f.endswith(".json")]
x = load_json(json_filepaths[0])
lookuptable = pd.DataFrame(x["look-up table"]).T
lookuptable.sort_values(by="mean")[["mean"]]
Out[10]:
| mean | |
|---|---|
| pelvis_start | 0.000 |
| femur_end | 13.616 |
| L5 | 25.532 |
| pelvis_end | 28.824 |
| L4 | 29.414 |
| L3 | 33.817 |
| kidney | 37.597 |
| L2 | 37.763 |
| L1 | 41.478 |
| lung_start | 44.143 |
| Th12 | 44.952 |
| Th11 | 47.725 |
| Th10 | 51.069 |
| Th9 | 53.994 |
| liver_end | 54.479 |
| Th8 | 56.856 |
| Th7 | 59.851 |
| Th6 | 63.177 |
| Th5 | 65.964 |
| Th4 | 68.499 |
| Th3 | 70.973 |
| Th2 | 73.401 |
| lung_end | 75.389 |
| Th1 | 75.794 |
| C7 | 77.937 |
| C6 | 79.497 |
| C5 | 81.826 |
| C4 | 83.817 |
| C3 | 85.647 |
| teeth | 86.908 |
| C2 | 87.206 |
| C1 | 89.234 |
| nose | 92.567 |
| eyes_end | 100.000 |
| head_end | 107.756 |
In [11]:
nifti_filepaths = [input_path + f.replace(".json", ".nii.gz") for f in df_cleaned.FILE]
# crop and save ct images to chest region
# crop_ct_images(nifti_filepaths, output_path, cropped_path, save=True)
# create body part meta data for cropped images
# bpreg_inference(cropped_path, cropped_json_path, plot=True, gpu_available=False)
10. Analyze preprocessed dataset¶
In [14]:
plot_volumes_interactive(cropped_path)
interactive(children=(BoundedFloatText(value=0.0, description='File:', max=108.0, step=1.0), Output()), _dom_c…
Thank you for your attention!¶
You find the notebook at: www.github.com/MIC-DKFZ/BodyPartRegression/docs/notebooks
Please feel free to reach out if you have questions.

In [ ]:
