


To get started: consult start
Sharing data features¶
Explore additional data¶
Once you analyse a corpus, it is likely that you produce data that others can reuse. Maybe you have defined a set of proper name occurrences, or special numerals, or you have computed part-of-speech assignments.
It is possible to turn these insights into new features, i.e. new .tf files with values assigned to specific nodes.
Make your own data¶
New data is a product of your own methods and computations in the first place. But how do you turn that data into new TF features? It turns out that the last step is not that difficult.
If you can shape your data as a mapping (dictionary) from node numbers (integers) to values (strings or integers), then TF can turn that data into a feature file for you with one command.
Share your new data¶
You can then easily share your new features on GitHub, so that your colleagues everywhere can try it out for themselves.
You can add such data on the fly, by passing a mod={org}/{repo}/{path} parameter,
or a bunch of them separated by commas.
If the data is there, it will be auto-downloaded and stored on your machine.
Let's do it.
%load_ext autoreload
%autoreload 2
import collections
import os
from tf.app import use
A = use("Nino-cunei/oldbabylonian", hoist=globals())
This is Text-Fabric 9.2.2 Api reference : https://annotation.github.io/text-fabric/tf/cheatsheet.html 67 features found and 0 ignored
Data: OLDBABYLONIAN, Character table, Feature docs
Features:
Old Babylonian Letters 1900-1600: Cuneiform tablets
Making data¶
We illustrate the data creation part by creating a new feature, ummama.
The idea is that we mark every sign reading that occurs between um-ma and ma some where in the first 3 lines of a face.
We want to mark every occurrence of such signs elsewhere in the corpus with ummama=1.
We only do it if the sign between the um-ma and ma (which must be on the same line) is not missing, damaged, or questionable.
The easiest way to get started is to run a query:
query = """
line ln<4
=: sign reading=um missing# damage# question#
<: sign reading=ma missing# damage# question#
% the next sign is the one that we are after
< sign missing# damage# question#
< sign reading=ma missing# damage# question#
"""
results = A.search(query)
1.29s 3466 results
A.table(results, end=10)
| n | p | line | sign | sign | sign | sign |
|---|---|---|---|---|---|---|
| 1 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | _{d} | ma |
| 2 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | en- | ma |
| 3 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | lil2_- | ma |
| 4 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | sza- | ma |
| 5 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | du- | ma |
| 6 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | u2- | ma |
| 7 | P509373 obverse:3 | um-ma _{d}en-lil2_-sza-du-u2-ni-ma | um- | ma | ni- | ma |
| 8 | P481190 obverse:3 | um-ma nu#-ur2#-i3-li2-szu-ma | um- | ma | i3- | ma |
| 9 | P481190 obverse:3 | um-ma nu#-ur2#-i3-li2-szu-ma | um- | ma | li2- | ma |
| 10 | P481190 obverse:3 | um-ma nu#-ur2#-i3-li2-szu-ma | um- | ma | szu- | ma |
Observe how the signs between um-ma and ma are picked up, except the damaged nu and ur2.
First we are collect these readings, and survey the frequencies in the result.
Some signs do not have a reading, but then they have a grapheme. If they do not have a grapheme, they might be comment signs, and we skip them.
umaReadings = collections.Counter()
# collect
for (line, um, ma1, sign, ma2) in results:
reading = F.reading.v(sign) or F.grapheme.v(sign)
if not reading:
continue
umaReadings[reading] += 1
# show
print(f"Found {len(umaReadings)} distinct readings")
limit = 20
for (reading, amount) in sorted(
umaReadings.items(),
key=lambda x: (-x[1], x[0]),
)[0:limit]:
print(f"{reading:<6} {amount:>4} x")
print(f" ... and {len(umaReadings) - limit} more ...")
Found 249 distinct readings d 324 x a 133 x ra 128 x mu 123 x am 112 x ha 99 x na 95 x pi2 94 x suen 78 x i 66 x ni 66 x szu 66 x utu 61 x um 59 x li2 55 x tum 55 x ma 50 x marduk 50 x bi 46 x nu 43 x ... and 229 more ...
Now we visit all signs in the whole corpus and check whether their reading or grapheme is in this set.
If so, we give that sign a value 1 in the dictionary ummama.
ummama = {}
allSigns = F.otype.s("sign")
for s in allSigns:
reading = F.reading.v(s) or F.grapheme.v(s)
if not reading:
continue
if reading in umaReadings:
ummama[s] = 1
print(f"Assigned `ummama=1` to {len(ummama)} sign occurrences out of {len(allSigns)}")
Assigned `ummama=1` to 182221 sign occurrences out of 203219
Note that the majority of all signs also occurs between um-ma and ma at the start of a document.
Maybe this is an indication that we are not capturing the idea of selecting specific signs, we may have to strengthen our search criterion.
But that is beyond this tutorial. We suppose these ummama words form a valuable set that we want to share.
Saving data¶
The documentation explains how to save this data into a text-fabric data file.
We choose a location where to save it, the exercises repository in the Nino-cunei organization, in the folder analysis.
In order to do this, we restart the TF API, but now with the desired output location in the locations parameter.
GITHUB = os.path.expanduser("~/github")
ORG = "Nino-cunei"
REPO = "exercises"
PATH = "bab-analysis"
VERSION = A.version
Note the version: we have built the version against a specific version of the data:
A.version
'1.0.6'
Later on, we pass this version on, so that users of our data will get the shared data in exactly the same version as their core data.
We have to specify a bit of metadata for this feature:
metaData = {
"ummama": dict(
valueType="int",
description="reading occurs somewhere between um-ma and ma",
creator="Dirk Roorda",
),
}
Now we can give the save command:
TF.save(
nodeFeatures=dict(ummama=ummama),
metaData=metaData,
location=f"{GITHUB}/{ORG}/{REPO}/{PATH}/tf",
module=VERSION,
)
0.00s Exporting 1 node and 0 edge and 0 config features to ~/github/Nino-cunei/exercises/bab-analysis/tf/1.0.6: | 0.15s T ummama to ~/github/Nino-cunei/exercises/bab-analysis/tf/1.0.6 0.16s Exported 1 node features and 0 edge features and 0 config features to ~/github/Nino-cunei/exercises/bab-analysis/tf/1.0.6
True
Sharing data¶
How to share your own data is explained in the documentation.
Here we show it step by step for the ummama feature.
If you commit your changes to the exercises repo, and have done a git push origin master,
you already have shared your data!
If you want to make a stable release, so that you can keep developing, while your users fall back on the stable data, you can make a new release.
Go to the GitHub website for that, go to your repo, and click Releases and follow the nudges.
If you want to make it even smoother for your users, you can zip the data and attach it as a binary to the release just created.
We need to zip the data in exactly the right directory structure. Text-Fabric can do that for us:
%%sh
text-fabric-zip Nino-cunei/exercises/bab-analysis/tf
This is a TF dataset Create release data for Nino-cunei/exercises/bab-analysis/tf Found 1 versions zip files end up in ~/Downloads/Nino-cunei-release/exercises zipping Nino-cunei/exercises 1.0.6 with 1 features ==> bab-analysis-tf-1.0.6.zip
All versions have been zipped, but it works OK if you only attach the newest version to the newest release.
If a user asks for an older version in this release, the system can still find it.
Here is the result for our case
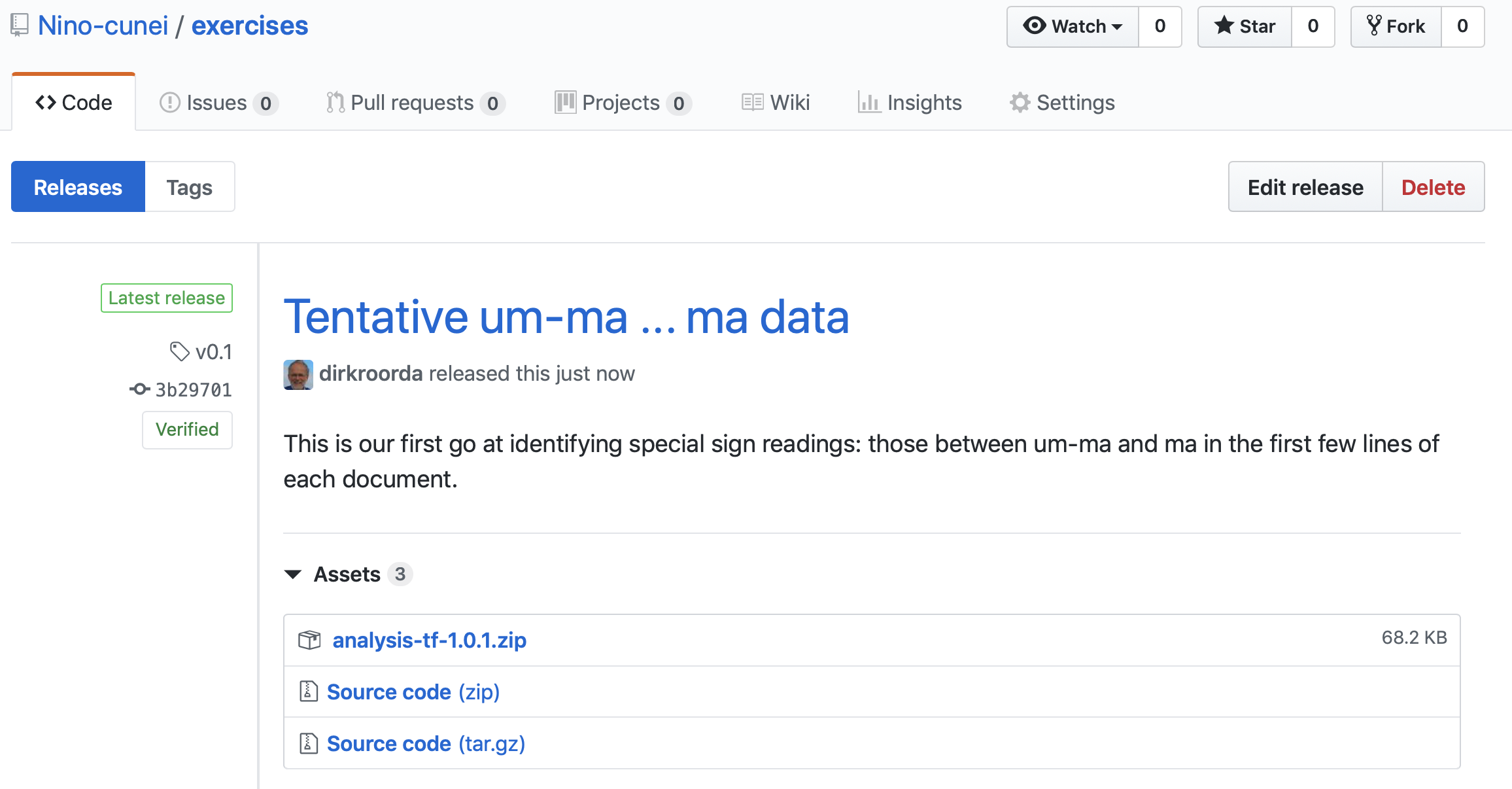
Use the data¶
We can use the data by calling it up when we say use('Nino-cunei/oldbabylonian', ...).
Here is how:
A = use(
"Nino-cunei/oldbabylonian:clone",
checkout="clone",
hoist=globals(),
mod="Nino-cunei/exercises/bab-analysis/tf:clone",
)
This is Text-Fabric 9.2.2 Api reference : https://annotation.github.io/text-fabric/tf/cheatsheet.html 68 features found and 0 ignored | | 0.34s C __characters__ from otext | 0.74s T ummama from ~/github/Nino-cunei/exercises/bab-analysis/tf/1.0.6
Data: OLDBABYLONIAN, Character table, Feature docs
Features:
Nino-cunei/exercises/bab-analysis/tf
Old Babylonian Letters 1900-1600: Cuneiform tablets
Above you see a new section in the feature list: Nino-cunei/exercises/analysis/tf with our foreign feature in it: ummama.
Now, suppose did not know much about this feature, then we would like to do a few basic checks:
F.ummama.freqList()
((1, 182221),)
We see that the feature has only one value, 1, and that 182222 nodes have it.
Which nodes have a ummama feature?
{F.otype.v(n) for n in N.walk() if F.ummama.v(n)}
{'sign'}
Only signs have the feature.
Let's have a look at a table of some ummama signs.
results = A.search(
"""
sign ummama
"""
)
0.23s 182221 results
A.table(results, start=1, end=20)
| n | p | sign |
|---|---|---|
| 1 | P509373 obverse:1 | [a- |
| 2 | P509373 obverse:1 | na] |
| 3 | P509373 obverse:1 | _{d} |
| 4 | P509373 obverse:1 | suen_- |
| 5 | P509373 obverse:1 | i- |
| 6 | P509373 obverse:1 | [din- |
| 7 | P509373 obverse:1 | nam] |
| 8 | P509373 obverse:2 | qi2- |
| 9 | P509373 obverse:2 | [ma] |
| 10 | P509373 obverse:3 | um- |
| 11 | P509373 obverse:3 | ma |
| 12 | P509373 obverse:3 | _{d} |
| 13 | P509373 obverse:3 | en- |
| 14 | P509373 obverse:3 | lil2_- |
| 15 | P509373 obverse:3 | sza- |
| 16 | P509373 obverse:3 | du- |
| 17 | P509373 obverse:3 | u2- |
| 18 | P509373 obverse:3 | ni- |
| 19 | P509373 obverse:3 | ma |
| 20 | P509373 obverse:4 | _{d} |
Now let's get some non-ummama signs:
results = A.search(
"""
sign ummama#
"""
)
0.12s 20998 results
A.table(results, start=1, end=20)
| n | p | sign |
|---|---|---|
| 1 | P509373 obverse:2 | bi2- |
| 2 | P509373 obverse:5 | t,u2- |
| 3 | P509373 obverse:6 | a2- |
| 4 | P509373 obverse:6 | gal2 |
| 5 | P509373 obverse:9 | 2(esze3) |
| 6 | P509373 obverse:9 | gud_ |
| 7 | P509373 obverse:10 | gar3_ |
| 8 | P509373 obverse:10 | ag- |
| 9 | P509373 obverse:10 | _uru_ |
| 10 | P509373 obverse:11 | kam_ |
| 11 | P509373 obverse:12 | _uru_ |
| 12 | P509373 obverse:12 | ak- |
| 13 | P509373 obverse:13 | 2(esze3) |
| 14 | P509373 obverse:13 | szuku_ |
| 15 | P509373 obverse:13 | _nagar- |
| 16 | P509373 obverse:14 | gar3 |
| 17 | P509373 obverse:14 | uru_ |
| 18 | P509373 obverse:14 | [...] |
| 19 | P509373 obverse:15 | [...] |
| 20 | P509373 obverse:$a | $ rest broken |
Let's get lines with both ummama and non-ummama signs:
results = A.search(
"""
line
sign ummama
sign ummama#
"""
)
0.58s 133413 results
A.table(results, start=1, end=2, condensed=True)
| n | p | line | sign | sign | sign | ||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | P509373 obverse:2 | qi2-bi2-[ma] | qi2- | bi2- | [ma] | ||||
| 2 | P509373 obverse:5 | li-ba-al-li-t,u2-u2-ka | li- | ba- | al- | li- | t,u2- | u2- | ka |
With highlights:
highlights = {}
for s in F.otype.s("sign"):
color = "lightsalmon" if F.ummama.v(s) else "mediumaquamarine"
highlights[s] = color
A.table(
results, start=1, end=10, baseTypes="sign", condensed=True, highlights=highlights
)
| n | p | line | sign | sign | sign | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | P509373 obverse:2 | qi2-bi2-[ma] | qi2- | bi2- | [ma] | ||||||||||||
| 2 | P509373 obverse:5 | li-ba-al-li-t,u2-u2-ka | li- | ba- | al- | li- | t,u2- | u2- | ka | ||||||||
| 3 | P509373 obverse:6 | {disz}sze-ep-_{d}suen a2-gal2 [dumu] um-mi-a-mesz_ | {disz} | sze- | ep- | _{d} | suen | a2- | gal2 | [dumu] | um- | mi- | a- | mesz_ | |||
| 4 | P509373 obverse:9 | 2(esze3) _a-sza3_ s,i-[bi]-it {disz}[ku]-un-zu-lum _sza3-gud_ | 2(esze3) | _a- | sza3_ | s,i- | [bi]- | it | {disz} | [ku]- | un- | zu- | lum | _sza3- | gud_ | ||
| 5 | P509373 obverse:10 | _a-sza3 a-gar3_ na-ag-[ma-lum] _uru_ x x x{ki} | [ma- | lum] | _uru_ | x | x | x | {ki} | _a- | sza3 | a- | gar3_ | na- | ag- | ||
| 6 | P509373 obverse:11 | sza _{d}utu_-ha-zi-[ir] isz-tu _mu 7(disz) kam_ id-di-nu-szum | sza | _{d} | utu_- | ha- | zi- | [ir] | isz- | tu | _mu | 7(disz) | kam_ | id- | di- | nu- | szum |
| 7 | P509373 obverse:12 | u3 i-na _uru_ x-szum{ki} sza-ak-nu id-di-a-am-ma | id- | di- | a- | am- | ma | u3 | i- | na | _uru_ | x- | szum | {ki} | sza- | ak- | nu |
| 8 | P509373 obverse:13 | 2(esze3) _a-sza3 szuku_ i-li-ib-bu s,i-bi-it _nagar-mesz_ | 2(esze3) | _a- | sza3 | szuku_ | i- | li- | ib- | bu | s,i- | bi- | it | _nagar- | mesz_ | ||
| 9 | P509373 obverse:14 | _a-sza3 a-gar3 uru_ ra-bu-um x [...] | _a- | sza3 | a- | gar3 | uru_ | ra- | bu- | um | x | [...] | |||||
| 10 | P509373 obverse:15 | x x x x x x [...] | x | x | [...] | x | x | x | x |
If we do a pretty display, the ummama feature shows up.
A.show(
results,
start=1,
end=3,
baseTypes="sign",
condensed=True,
withNodes=True,
highlights=highlights,
)
line 1
line 2
line 3
Or in the context of a whole face:
A.show(
results,
start=1,
end=1,
condensed=True,
condenseType="face",
withNodes=False,
highlights=highlights,
)
face 1
All together!¶
If more researchers have shared data modules, you can draw them all in.
Then you can design queries that use features from all these different sources.
In that way, you build your own research on top of the work of others.
Hover over the features to see where they come from, and you'll see they come from your local GitHub repo.
All chapters:
- start become an expert in creating pretty displays of your text structures
- display become an expert in creating pretty displays of your text structures
- search turbo charge your hand-coding with search templates
- exportExcel make tailor-made spreadsheets out of your results
- share draw in other people's data and let them use yours
- similarLines spot the similarities between lines
See the cookbook for recipes for small, concrete tasks.
CC-BY Dirk Roorda