Parsl: Workflow Tutorial¶
In this tutorial, you will be able to first try a few Parsl dataflows (examples 1-4) on your local machine, to get a sense of the library. Then, in examples 5-7 you will run similar dataflows on any resource you may have access to, such as clouds (Amazon Web Services) and HPC systems, to see how more complex workflows can be expressed with Parsl.
Tutorial Section One¶
This section will provide a walk-through for getting a simple "mock" science application running using Parsl on your local machine. | Note: the mock science apps are included in the apps directory of the tutorial repository.
Example 1: Run a single application using Parsl¶
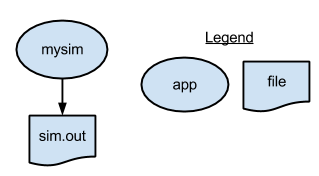
The first Parsl script runs simulate.sh to generate a single random number. It writes the number to stdout. You can run the following script repeatedly to see different "simulated" results.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
parsl.load(config)
@bash_app
def mysim(stdout="output/p1.out", stderr="output/p1.err"):
"""Set this example up as a bash app by returning the
command line app to be called, in this case simulate"""
return "app/simulate"
# call the mysim app and wait for the result
mysim().result()
# open the output file and read the result
with open('output/p1.out', 'r') as f:
print(f.read())
parsl.clear()
Debugging¶
Debugging parallel dataflows is often more complicated than with serial programs. The easiest way to debug a Parsl application is via Python loggers. The following example shows how to enable logging. In this case we log the debug stream to the console. You can also log to a file using the set_file_logger function.
When trying to debug individual apps it is often best to first capture stdout and stderr to files. These files will capture any text output by the app which may indicate app behavior.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
from parsl import set_stream_logger, NullHandler
# set the stream logger to print debug messages
set_stream_logger()
parsl.load(config)
@bash_app
def mysim(stdout="output/p1-debug.out", stderr="output/p1-debug.err"):
"""Set this example up as a bash app by returning the
command line app to be called, in this case simulate"""
return "app/simulate"
mysim().result()
with open('output/p1-debug.out', 'r') as f:
print(f.read())
parsl.clear()
Example 2: Run a single Python function using Parsl¶
The second example mirrors the first, however instead of using an external application it uses a Python function to simulate a science app again by generating a random number. In this case the Python app returns the simulated value as a Python object rather than using an external file.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
parsl.load(config)
@python_app
def mysim():
from random import randint
"""Generate a random integer and return it"""
return randint(1,100)
print(mysim().result())
parsl.clear()
Example 3: Running an ensemble of many apps in parallel with a loop¶
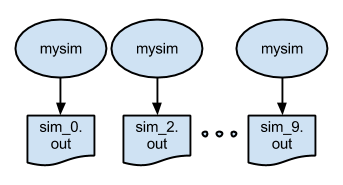
The third example shows how Parsl can be used to run many simulations in parallel. In this case, we define the same Parsl simulation app (simulate). The Python script then loops calling this app. Parsl ensures that each independent instance of the app will execute in parallel. Note: rather than use stdout the simulation app redirects the output to a specified file.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
from parsl.data_provider.files import File
parsl.load(config)
@bash_app
def mysim(outputs=[], stdout="output/p3.out", stderr="output/p3.err"):
"""Call simulate and return results in the output file"""
return 'app/simulate > {0}'.format(outputs[0].filepath)
# loop to execute the simulation app 5 times
results = []
for i in range(5):
out_file = "output/p3_sim_{0}".format(i)
results.append(mysim(outputs=[File(out_file)]))
# print each job status, initially all are running
print ("Job Status: {}".format([r.done() for r in results]))
# wait for all apps to complete
[r.result() for r in results]
# print each job status, they will now be finished
print ("Job Status: {}".format([r.done() for r in results]))
# collect up the output files and print their values
outputs = [r.outputs[0] for r in results]
for o in outputs:
with open(o.filename, 'r') as f:
print(f.read().strip())
parsl.clear()
Example 4: Analyzing results of a parallel ensemble¶
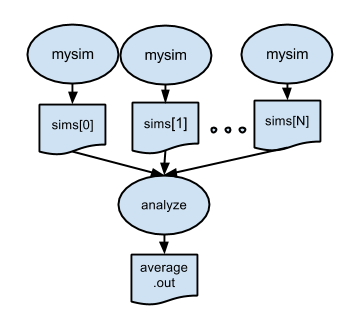
After all the parallel simulations in an ensemble run have completed, it is typically necessary to gather and analyze their results with some kind of post-processing analysis program or script. The fourth example introduces such a postprocessing step. In this case, the files created by all of the parallel runs of simulate will be averaged by the "analysis application" stats.
Note: in this example we do not block on the outputs of the simulation app, rather the futures are passed directly to the analysis application. Parsl will manage these dependencies and only execute the analysis app when the simulation app completes.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
from parsl.data_provider.files import File
parsl.load(config)
@bash_app
def mysim(outputs=[],
stdout="output/p4_sim.out",
stderr="output/p4_sim.err"):
"""Call simulate and return results in the output file"""
return 'app/simulate > {0}'.format(outputs[0].filepath)
@bash_app
def stats(inputs=[],
outputs=[],
stderr='output/p4_stats.err',
stdout='output/p4_stats.out'):
"""call stats with all simulation results as inputs"""
return "app/stats {0} > {1}".format(" ".join(i.filepath for i in inputs), outputs[0].filepath)
# call the simulation app 5 times
results = []
for i in range(5):
out_file = "output/p4_sim_{0}".format(i)
results.append(mysim(outputs=[File(out_file)]))
# collect the output data futures
sim_outputs = [r.outputs[0] for r in results]
# run the stats app
s = stats(inputs=sim_outputs, outputs=[File("output/p4_stats.txt")])
# wait for the result
s.result()
# print the result
with open('output/p4_stats.txt', 'r') as f:
print(f.read())
parsl.clear()
Tutorial Section Two¶
This section introduces execution of Parsl scripts on remote computational resources.
Parsl supports a variety of resource providers as well as methods for submitting workload to those resources (e.g., pilot jobs). Example configurations for compute resources such as UChicago Midway, NERSC Cori, NCSA BlueWaters, and ANL Cooley are included in the config directory of the tutorial repository.
A Parsl script can be executed from a login node by using these configuration files directly. Parsl also supports a more advanced submission model in which a SSH channel is used to submit wokloads from an external machine (e.g., your laptop). In this case the machine on which the script is executed must be accessible from the cluster (i.e., you will need to ensure that firewall rules allow pilot job connections back to your laptop).
Example 5: Running a simple app using pilot jobs¶
In this example we show how one might develop a script to run on a remote resource. We first develop a Parsl script for sorting a file. We initially run this script using the local ThreadPoolExecutor on a laptop, we subsequently extend this example to submit the job via the HighThroughputExecutor pilot job model locally and then on a remote resource. We finally show how Parsl's SSH channel can be used to connect to the remote resource directly from your laptop.
This script uses the sort application to sort the numbers from an unsorted file.
Example 5.a: Running the sort app locally using threads¶
First we use the local thread executor to run the sort command locally.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.local_threads import config
from parsl.data_provider.files import File
parsl.load(config)
@bash_app
def sort(unsorted, outputs=[]):
"""Call sort executable on the input file"""
return "sort -g {} > {}".format(unsorted.filepath, outputs[0].filepath)
# call the sort app on the unsorted.txt file
# save the results to a_sorted.txt
s = sort(File("input/unsorted.txt"), outputs=[File("output/a_sorted.txt")])
# wait for the result
output_file = s.outputs[0].result()
# print the contents of the unsorted and sorted files
print("Contents of the unsorted.txt file:")
with open('input/unsorted.txt', 'r') as f:
print(f.read().replace("\n",","))
print("\nContents of the sorted output file:")
with open(output_file, 'r') as f:
print(f.read().replace("\n",","))
parsl.clear()
Example 5.b: Running the sort app locally using pilot jobs¶
We now use IPyParallel to run the sort command using a pilot job model. In this case we use the htex_local configuration to tell Parsl to use local HighThroughputExecutor.
You will notice that apart from the configuration and instantiation of the DataFlowKernal this script is identical to the previous script.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.configs.htex_local import config
from parsl.data_provider.files import File
parsl.load(config)
@bash_app
def sort(unsorted, outputs=[]):
"""Call sort executable on the input file"""
return "sort -g {} > {}".format(unsorted.filepath, outputs[0].filepath)
s = sort(File("input/unsorted.txt"), outputs=[File("output/b_sorted.txt")])
output_file = s.outputs[0].result()
print("Contents of the unsorted.txt file:")
with open('input/unsorted.txt', 'r') as f:
print(f.read().replace("\n",","))
print("\nContents of the sorted output file:")
with open(output_file, 'r') as f:
print(f.read().replace("\n",","))
parsl.clear()
Example 5.c: Running the sort app on a cluster using pilot jobs¶
We now take the previous example and run it on a cluster using HighThroughputExecutor. To run this example you will need to execute the script from a login node and uncomment the configuration needed for your cluster.
The following example uses a basic configuration for TACC's Frontera. You will need to update the scheduler_options with your allocation and the conda environment specified by worker_init.
Note: you will need to either install the Parsl library on that login node or use an existing environment. You will also need to download the parsl-tutorial repository and ensure that it is avaialble on the worker nodes via a shared file system.
import parsl
from parsl.app.app import python_app, bash_app
from parsl.data_provider.files import File
import os
from parsl.config import Config
from parsl.channels import LocalChannel
from parsl.providers import SlurmProvider
from parsl.executors import HighThroughputExecutor
from parsl.launchers import SrunLauncher
from parsl.addresses import address_by_hostname
""" This config assumes that it is used to launch parsl tasks from the login nodes
of Frontera at TACC. Each job submitted to the scheduler will request 2 nodes for 10 minutes.
"""
config = Config(
executors=[
HighThroughputExecutor(
label="frontera_htex",
address=address_by_hostname(),
max_workers=56,
provider=SlurmProvider(
cmd_timeout=60,
channel=LocalChannel(),
nodes_per_block=1,
partition='development',
scheduler_options='#SBATCH -A ALLOCATION', # Enter allocation
worker_init='conda activate CONDA_ENV', # Enter conda environment
walltime='00:10:00',
launcher=SrunLauncher(),
),
)
],
)
parsl.load(config)
@bash_app
def sort(unsorted, outputs=[]):
"""Call sort executable on the input file"""
return "sleep 30; sort -g {} > {}".format(unsorted.filepath, outputs[0].filepath)
s = sort(File(os.path.abspath("input/unsorted.txt")),
outputs=[File(os.path.abspath("output/sorted_c.txt"))])
output_file = s.outputs[0].result()
print("Contents of the unsorted.txt file:")
with open('input/unsorted.txt', 'r') as f:
print(f.read().replace("\n",","))
print("\nContents of the sorted output file:")
with open(output_file, 'r') as f:
print(f.read().replace("\n",","))
parsl.clear()
Example 6: MPI Hello¶
The final example is a basic "Hello World!" example that shows you how to run MPI applications. Here we have a simple MPI code mpi_hello.c that has each MPI rank sleep for a user-specified duration and then print the processor name on which the rank is executing followed by "Hello World!".
The following script is designed to be run from the login node. Like the previous examples you will need to update the shared directory. You may optionally use an SSH channel by following the previous instructions. If running from the login node you will not need the remote configuration details.
Given the range of MPI libraries installed on each site you will need to specify the site-specific MPI compiler (e.g., mpicc, cc) and the way to execute the MPI job (e.g., mpirun, srun).
import parsl
from parsl.app.app import python_app, bash_app
# Update to import config for your machine
from config.midway import config
import os
parsl.load(config)
remote = False
shared_dir = 'SHARED_DIRECTORY' # path to the tutorial repository
@bash_app
def compile_app(dirpath, stdout=None, stderr=None, compiler="mpicc"):
"""Compile mpi app using site-specific compiler.
E.g., midway compiler = mpicc, Cori compiler= cc
"""
return '''cd {0}; make clean; make CC={1} '''.format(dirpath, compiler)
@bash_app
def mpi_hello(dirpath, launcher="mpirun", app="mpi_hello", nproc=20, outputs=[]):
"""Call compiled mpi executable with local mpilib.
Works natively for openmpi mpiexec, mpirun, orterun, oshrun, shmerun
mpiexec is default"""
import os
if launcher == "mpirun" :
return "mpirun -np {} {} &> {};".format(nproc, os.path.join(dirpath,app), outputs[0])
elif launcher == "srun" :
return "srun -n {} ./{} &> {};".format(nproc, os.path.join(dirpath,app), outputs[0])
# complile the app and wait for it to complete (.result())
compile_app(dirpath=os.path.join(shared_dir, "mpi_apps"),
stdout=os.path.join(shared_dir, "mpi_apps.compile.out"),
stderr=os.path.join(shared_dir, "mpi_apps.compile.err",),
compiler='mpicc'
).result()
# run the mpi app
hello = mpi_hello(os.path.join(shared_dir, "mpi_apps"),
launcher="mpirun",
outputs=[os.path.join(shared_dir, "mpi_apps", "hello.txt")])
output_file = hello.outputs[0].result()
# if running remotely using SSH, copy the file back to the host
if remote:
dfk.executor.execution_provider.channel.pull_file(output_file, '.')
# read the result file
with open(output_file, 'r') as f:
print(f.read())
parsl.clear()