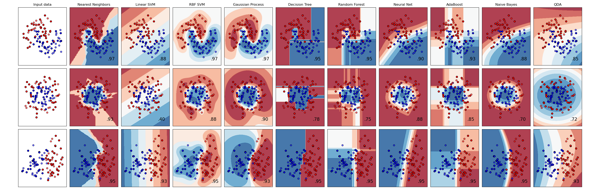
Scikit Learn Machine Learning Library¶

Dimensionality reduction¶
*** Reducing the number of random variables to consider.
- Applications: Visualization, Increased efficiency
- Algorithms: PCA, feature selection, non-negative matrix factorization
from sklearn import datasets
iris = datasets.load_iris()
type(iris)
sklearn.utils.Bunch
print(iris.data)
[[5.1 3.5 1.4 0.2] [4.9 3. 1.4 0.2] [4.7 3.2 1.3 0.2] [4.6 3.1 1.5 0.2] [5. 3.6 1.4 0.2] [5.4 3.9 1.7 0.4] [4.6 3.4 1.4 0.3] [5. 3.4 1.5 0.2] [4.4 2.9 1.4 0.2] [4.9 3.1 1.5 0.1] [5.4 3.7 1.5 0.2] [4.8 3.4 1.6 0.2] [4.8 3. 1.4 0.1] [4.3 3. 1.1 0.1] [5.8 4. 1.2 0.2] [5.7 4.4 1.5 0.4] [5.4 3.9 1.3 0.4] [5.1 3.5 1.4 0.3] [5.7 3.8 1.7 0.3] [5.1 3.8 1.5 0.3] [5.4 3.4 1.7 0.2] [5.1 3.7 1.5 0.4] [4.6 3.6 1. 0.2] [5.1 3.3 1.7 0.5] [4.8 3.4 1.9 0.2] [5. 3. 1.6 0.2] [5. 3.4 1.6 0.4] [5.2 3.5 1.5 0.2] [5.2 3.4 1.4 0.2] [4.7 3.2 1.6 0.2] [4.8 3.1 1.6 0.2] [5.4 3.4 1.5 0.4] [5.2 4.1 1.5 0.1] [5.5 4.2 1.4 0.2] [4.9 3.1 1.5 0.1] [5. 3.2 1.2 0.2] [5.5 3.5 1.3 0.2] [4.9 3.1 1.5 0.1] [4.4 3. 1.3 0.2] [5.1 3.4 1.5 0.2] [5. 3.5 1.3 0.3] [4.5 2.3 1.3 0.3] [4.4 3.2 1.3 0.2] [5. 3.5 1.6 0.6] [5.1 3.8 1.9 0.4] [4.8 3. 1.4 0.3] [5.1 3.8 1.6 0.2] [4.6 3.2 1.4 0.2] [5.3 3.7 1.5 0.2] [5. 3.3 1.4 0.2] [7. 3.2 4.7 1.4] [6.4 3.2 4.5 1.5] [6.9 3.1 4.9 1.5] [5.5 2.3 4. 1.3] [6.5 2.8 4.6 1.5] [5.7 2.8 4.5 1.3] [6.3 3.3 4.7 1.6] [4.9 2.4 3.3 1. ] [6.6 2.9 4.6 1.3] [5.2 2.7 3.9 1.4] [5. 2. 3.5 1. ] [5.9 3. 4.2 1.5] [6. 2.2 4. 1. ] [6.1 2.9 4.7 1.4] [5.6 2.9 3.6 1.3] [6.7 3.1 4.4 1.4] [5.6 3. 4.5 1.5] [5.8 2.7 4.1 1. ] [6.2 2.2 4.5 1.5] [5.6 2.5 3.9 1.1] [5.9 3.2 4.8 1.8] [6.1 2.8 4. 1.3] [6.3 2.5 4.9 1.5] [6.1 2.8 4.7 1.2] [6.4 2.9 4.3 1.3] [6.6 3. 4.4 1.4] [6.8 2.8 4.8 1.4] [6.7 3. 5. 1.7] [6. 2.9 4.5 1.5] [5.7 2.6 3.5 1. ] [5.5 2.4 3.8 1.1] [5.5 2.4 3.7 1. ] [5.8 2.7 3.9 1.2] [6. 2.7 5.1 1.6] [5.4 3. 4.5 1.5] [6. 3.4 4.5 1.6] [6.7 3.1 4.7 1.5] [6.3 2.3 4.4 1.3] [5.6 3. 4.1 1.3] [5.5 2.5 4. 1.3] [5.5 2.6 4.4 1.2] [6.1 3. 4.6 1.4] [5.8 2.6 4. 1.2] [5. 2.3 3.3 1. ] [5.6 2.7 4.2 1.3] [5.7 3. 4.2 1.2] [5.7 2.9 4.2 1.3] [6.2 2.9 4.3 1.3] [5.1 2.5 3. 1.1] [5.7 2.8 4.1 1.3] [6.3 3.3 6. 2.5] [5.8 2.7 5.1 1.9] [7.1 3. 5.9 2.1] [6.3 2.9 5.6 1.8] [6.5 3. 5.8 2.2] [7.6 3. 6.6 2.1] [4.9 2.5 4.5 1.7] [7.3 2.9 6.3 1.8] [6.7 2.5 5.8 1.8] [7.2 3.6 6.1 2.5] [6.5 3.2 5.1 2. ] [6.4 2.7 5.3 1.9] [6.8 3. 5.5 2.1] [5.7 2.5 5. 2. ] [5.8 2.8 5.1 2.4] [6.4 3.2 5.3 2.3] [6.5 3. 5.5 1.8] [7.7 3.8 6.7 2.2] [7.7 2.6 6.9 2.3] [6. 2.2 5. 1.5] [6.9 3.2 5.7 2.3] [5.6 2.8 4.9 2. ] [7.7 2.8 6.7 2. ] [6.3 2.7 4.9 1.8] [6.7 3.3 5.7 2.1] [7.2 3.2 6. 1.8] [6.2 2.8 4.8 1.8] [6.1 3. 4.9 1.8] [6.4 2.8 5.6 2.1] [7.2 3. 5.8 1.6] [7.4 2.8 6.1 1.9] [7.9 3.8 6.4 2. ] [6.4 2.8 5.6 2.2] [6.3 2.8 5.1 1.5] [6.1 2.6 5.6 1.4] [7.7 3. 6.1 2.3] [6.3 3.4 5.6 2.4] [6.4 3.1 5.5 1.8] [6. 3. 4.8 1.8] [6.9 3.1 5.4 2.1] [6.7 3.1 5.6 2.4] [6.9 3.1 5.1 2.3] [5.8 2.7 5.1 1.9] [6.8 3.2 5.9 2.3] [6.7 3.3 5.7 2.5] [6.7 3. 5.2 2.3] [6.3 2.5 5. 1.9] [6.5 3. 5.2 2. ] [6.2 3.4 5.4 2.3] [5.9 3. 5.1 1.8]]
type(iris.data)
numpy.ndarray
Loading from external datasets¶
http://scikit-learn.org/stable/datasets/index.html#external-datasets
from sklearn import svm
clf = svm.SVC()
X, y = iris.data, iris.target
clf.fit(X, y)
SVC(C=1.0, cache_size=200, class_weight=None, coef0=0.0, decision_function_shape='ovr', degree=3, gamma='auto', kernel='rbf', max_iter=-1, probability=False, random_state=None, shrinking=True, tol=0.001, verbose=False)
len(iris.data)
150
# Persistent Models using pickle
import pickle
s = pickle.dumps(clf)
clf2 = pickle.loads(s)
clf2.predict(X[0:1])
array([0])
type(clf2)
sklearn.svm.classes.SVC
y[0]
0
Big data pickle¶
In the specific case of the scikit, it may be more interesting to use joblib’s replacement of pickle (joblib.dump & joblib.load), which is more efficient on big data, but can only pickle to the disk and not to a string:
from sklearn.externals import joblib
joblib.dump(clf, 'filename.pkl')
Full Scikit learn process¶
- Define Problem.
- Prepare Data.
- Evaluate Algorithms.
- Improve Results.
- Present Results.
from https://machinelearningmastery.com/machine-learning-in-python-step-by-step/
# Library setup/import
import pandas as pd
from pandas.plotting import scatter_matrix
import matplotlib.pyplot as plt
from sklearn import model_selection
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
# Load dataset
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data" #UCI ml archive
names = ['sepal-length', 'sepal-width', 'petal-length', 'petal-width', 'class']
dataset = pd.read_csv(url, names=names)
# Summarizing data
# shape
print(dataset.shape)
(150, 5)
dataset.head()
| sepal-length | sepal-width | petal-length | petal-width | class | |
|---|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 | Iris-setosa |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 | Iris-setosa |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 | Iris-setosa |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 | Iris-setosa |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 | Iris-setosa |
# Stat summary
dataset.describe()
| sepal-length | sepal-width | petal-length | petal-width | |
|---|---|---|---|---|
| count | 150.000000 | 150.000000 | 150.000000 | 150.000000 |
| mean | 5.843333 | 3.054000 | 3.758667 | 1.198667 |
| std | 0.828066 | 0.433594 | 1.764420 | 0.763161 |
| min | 4.300000 | 2.000000 | 1.000000 | 0.100000 |
| 25% | 5.100000 | 2.800000 | 1.600000 | 0.300000 |
| 50% | 5.800000 | 3.000000 | 4.350000 | 1.300000 |
| 75% | 6.400000 | 3.300000 | 5.100000 | 1.800000 |
| max | 7.900000 | 4.400000 | 6.900000 | 2.500000 |
# Class Distribution
print(dataset.groupby('class').size()) # notice aggregration similar to SQL
class Iris-setosa 50 Iris-versicolor 50 Iris-virginica 50 dtype: int64
# Visualization
# univariate plots, that is, plots of each individual variable.
# box and whisker plots
dataset.plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False)
plt.show()
# histograms
dataset.hist()
plt.show()
# Multivariate Plots
# scatter plot matrix
scatter_matrix(dataset)
plt.show()
# Note the diagonal grouping of some pairs of attributes. This suggests a high correlation and a predictable relationship.
Algorithm Evaluation¶
Create a Validation Dataset¶
# Split-out validation dataset
# We will split the loaded dataset into two,
# 80% of which we will use to train our models and
# 20% that we will hold back as a validation dataset.
array = dataset.values
X = array[:,0:4] # our attributes
Y = array[:,4] # our Class
validation_size = 0.20
seed = 7
X_train, X_validation, Y_train, Y_validation = model_selection.train_test_split(X, Y, test_size=validation_size, random_state=seed)
# WE have now have training data in the X_train and Y_train for preparing models and a X_validation and Y_validation sets that we can use later.
Test Harness¶
We will use 10-fold cross validation to estimate accuracy.
# Test options and evaluation metric
seed = 7 # just to have a consistent pseudo random numers, could be any constant
scoring = 'accuracy'
Build Models¶
Let’s evaluate 6 different algorithms:¶
- Logistic Regression (LR)
- Linear Discriminant Analysis (LDA)
- K-Nearest Neighbors (KNN).
- Classification and Regression Trees (CART).
- Gaussian Naive Bayes (NB).
- Support Vector Machines (SVM)
# Spot Check Algorithms
models = []
models.append(('LR', LogisticRegression()))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier()))
models.append(('NB', GaussianNB()))
models.append(('SVM', SVC()))
# evaluate each model in turn
results = []
names = []
for name, model in models:
kfold = model_selection.KFold(n_splits=10, random_state=seed)
cv_results = model_selection.cross_val_score(model, X_train, Y_train, cv=kfold, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
LR: 0.966667 (0.040825) LDA: 0.975000 (0.038188) KNN: 0.983333 (0.033333) CART: 0.966667 (0.040825) NB: 0.975000 (0.053359) SVM: 0.991667 (0.025000)
# plot comparison
# Compare Algorithms
fig = plt.figure()
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
Make Predictions¶
The KNN algorithm was the most accurate model that we tested. Now we want to get an idea of the accuracy of the model on our validation set.
This will give us an independent final check on the accuracy of the best model. It is valuable to keep a validation set just in case you made a slip during training, such as overfitting to the training set or a data leak. Both will result in an overly optimistic result.
We can run the KNN model directly on the validation set and summarize the results as a final accuracy score, a confusion matrix and a classification report.
# Make predictions on validation dataset
knn = KNeighborsClassifier()
knn.fit(X_train, Y_train)
predictions = knn.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
0.9
[[ 7 0 0]
[ 0 11 1]
[ 0 2 9]]
precision recall f1-score support
Iris-setosa 1.00 1.00 1.00 7
Iris-versicolor 0.85 0.92 0.88 12
Iris-virginica 0.90 0.82 0.86 11
avg / total 0.90 0.90 0.90 30
We can see that the accuracy is 0.9 or 90%. The confusion matrix provides an indication of the three errors made. Finally, the classification report provides a breakdown of each class by precision, recall, f1-score and support showing excellent results (granted the validation dataset was small).