LSPR sensor response of silver NP to BSA¶
In the section 03_LSPR_sensor_response_POC we presented a proof of concept of LSPR sensor response to BSA. LSPR biosensors detect a target molecule by monitoring plasmon resonance frequency changes due to the proximity of the target to the metallic nanoparticle.
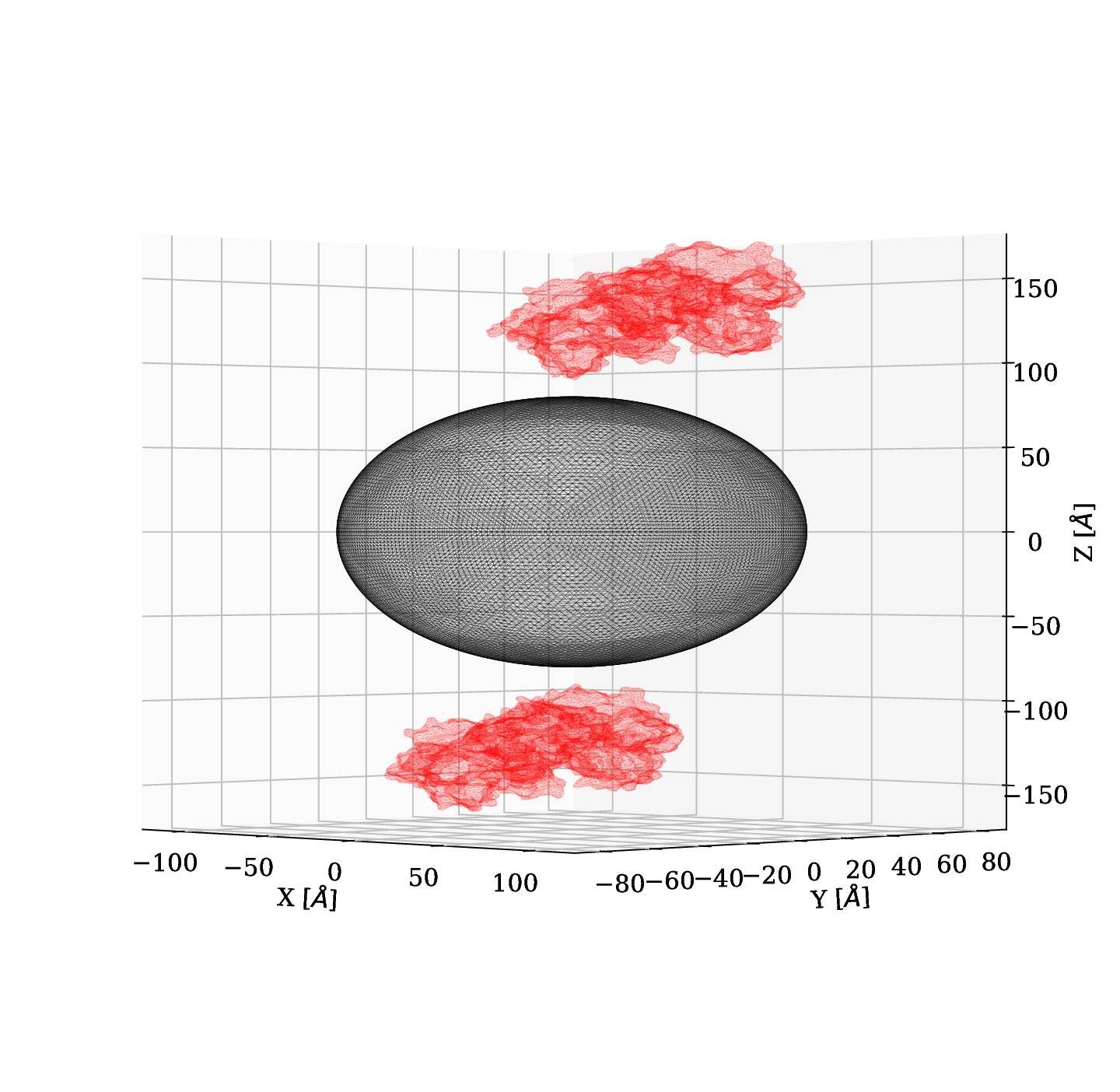
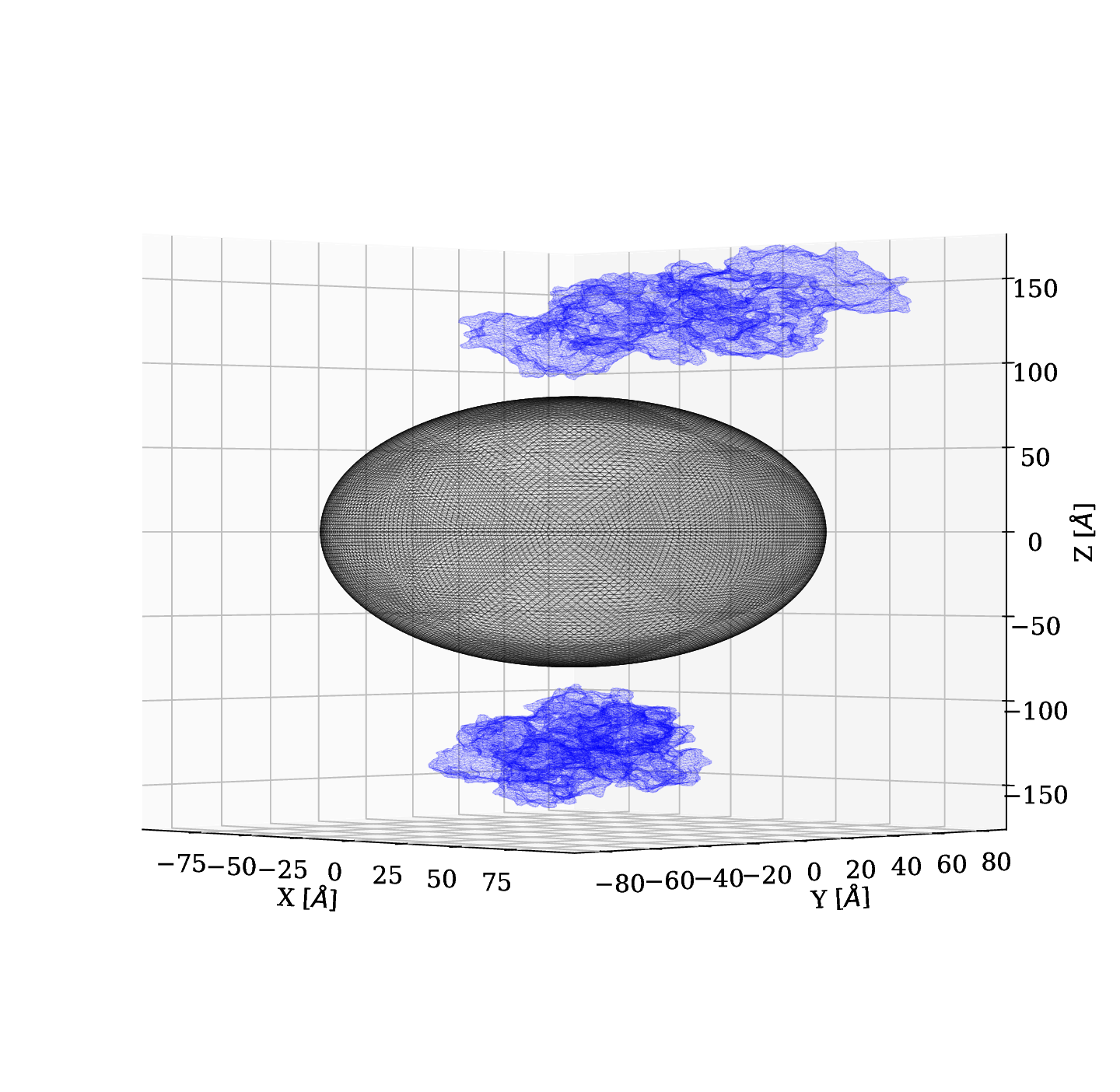
In the POC study, we used spheres to represent BSA. In this section we present results obtained by using a mesh of the crystal structure of Bovine Serum Albumina and its charge distribution.
Initially, we used a silver nanoparticle of $r=25 \; nm$ and we set one BSA protein at a distance of $1 \; nm$ along the z-axis and we had an electric field of -1 $[e]/([{\unicode{x212B}^2}][\epsilon_0])$ (equivalent to $-1.8095 \; \text{x} \; 10^{12} \;[V]/[m]$), where the minus indicates the electric field is pointing in -z direction. For this case we didn't obeserve significant changes.
After consulting parameters picked for similar studies in Pahn, et al. (1) and Tang, et al. (2), we picked a silver nanoparticle $r=8 \; nm$ and an electric field of $-6.64 \; \text{x} \; 10^{9} \;[V]/[m]$ (units of the code it's $\sim -0.00367 [e]/([{\unicode{x212B}^2}][\epsilon_0])$).
Preliminary conclusions:
- The intensity of the electric field affects only when we include the charges.
We noticed that if we remove the charges from inside the protein and we vary the magnitud and direction of the electric field, the results obtained do not change.
- The electric field affects the magnitud of the peak.
We observed that when charges are present, the magnitud of the peak varies depending on the electric field.
- When we tilt the angle, we don't detect changes.
The shift amplitud and the magnitud of the peak is preserved.
- The orientation seems to have an effect on the shift when we rotate the protein.
It is still note clear what it causes this effect. We noticed that the charge that is closer to the surface varies when we rotate but not when we tilt, this might be a possible explanantion.
- The location of the protein affects the shift of the peak.
After multiple simulations with electric field in -z-axis, we realized that two proteins located at 1 nm +/- z-axis with its dipole moment aligned with the y-axis, influence more than those proteins located at 1 nm +/- y-axis or x-axis.
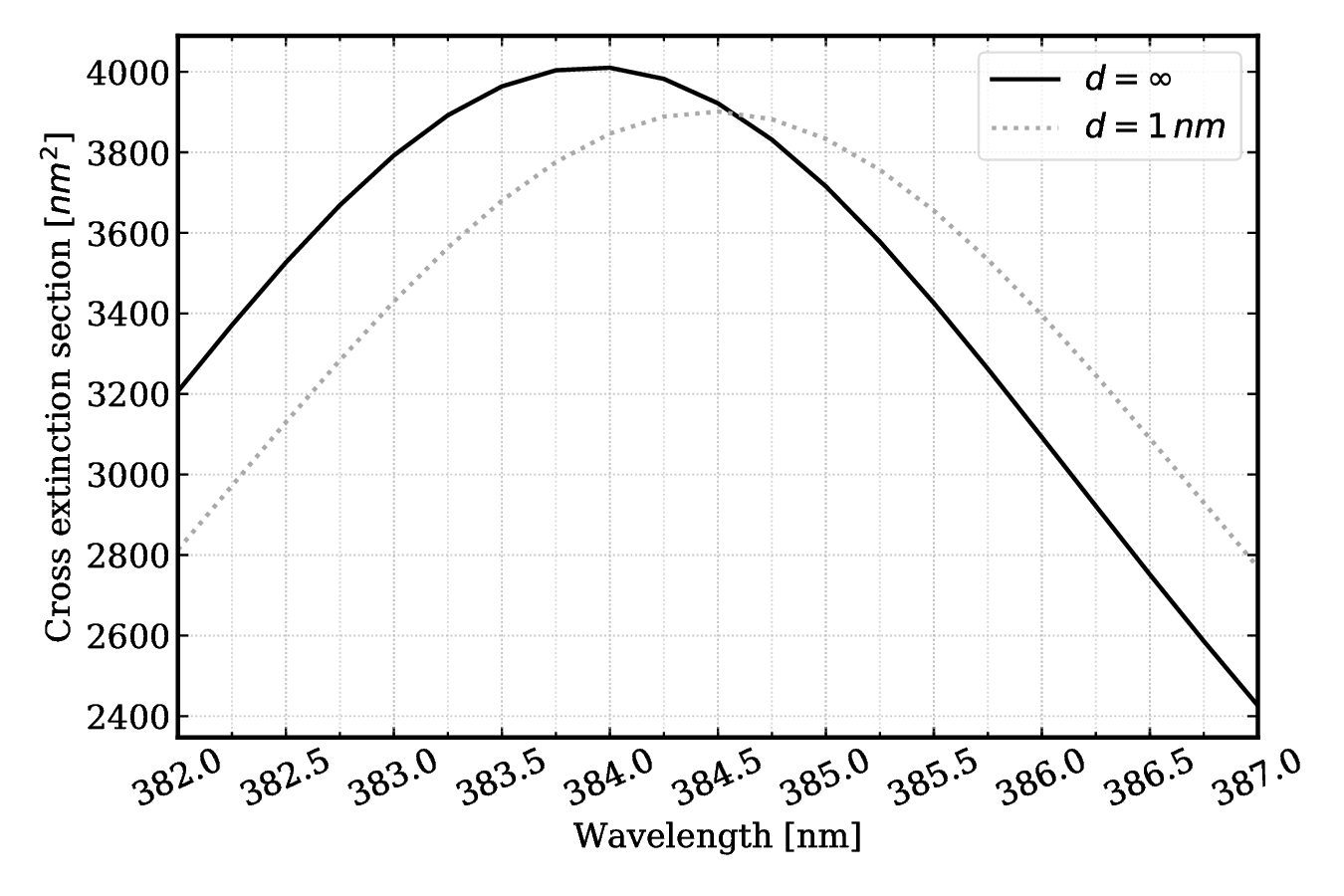
#### Fig 2. Close-up of spectrum shift d = 1 nm.
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3901$ and it occurs at a wavelength of 384.5 nm
Shift: 0.5 nm

#### Fig 3. Close-up of spectrum shift d = 0.5 nm.
Cext max at $d=\infty$ is $\sim 4010 \, nm^2$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=0.5$ nm is $\sim 3843 \, nm^2$ and it occurs at a wavelength of 384.75 nm
Shift: 0.75 nm
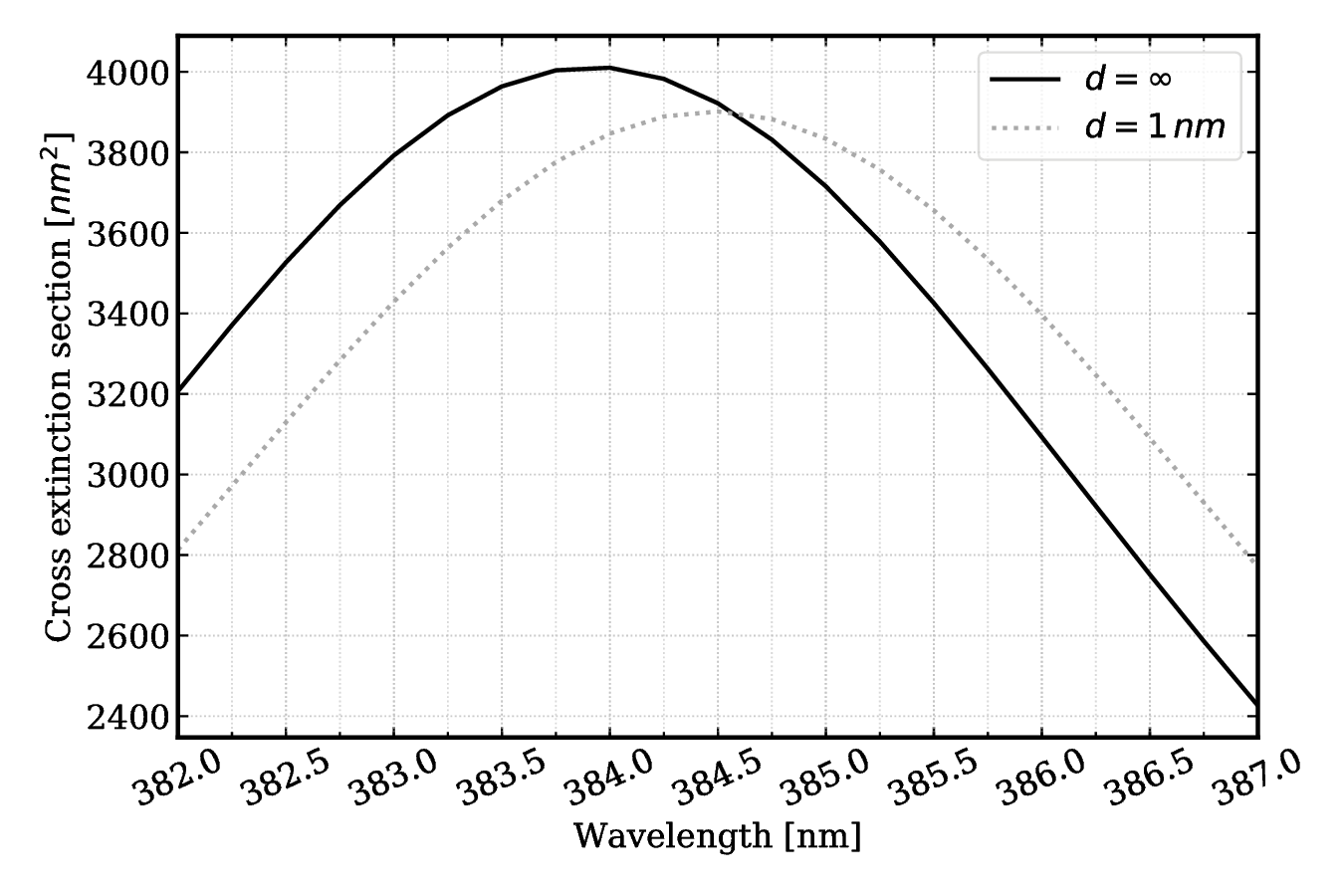
#### Fig 5. Close-up of spectrum shift d = 1 nm with protein tilted 30 deg.
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3901$ and it occurs at a wavelength of 384.5 nm
Shift: 0.5 nm
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3903$ and it occurs at a wavelength of 384.5 nm
Shift: 0.5 nm
Results of 2 BSA located at 1 nm of +/- z with a tilt angle of 45 deg¶
We tested the case of tilt 45 deg and we obtained similar results. No changes in the magnitud of the shift.
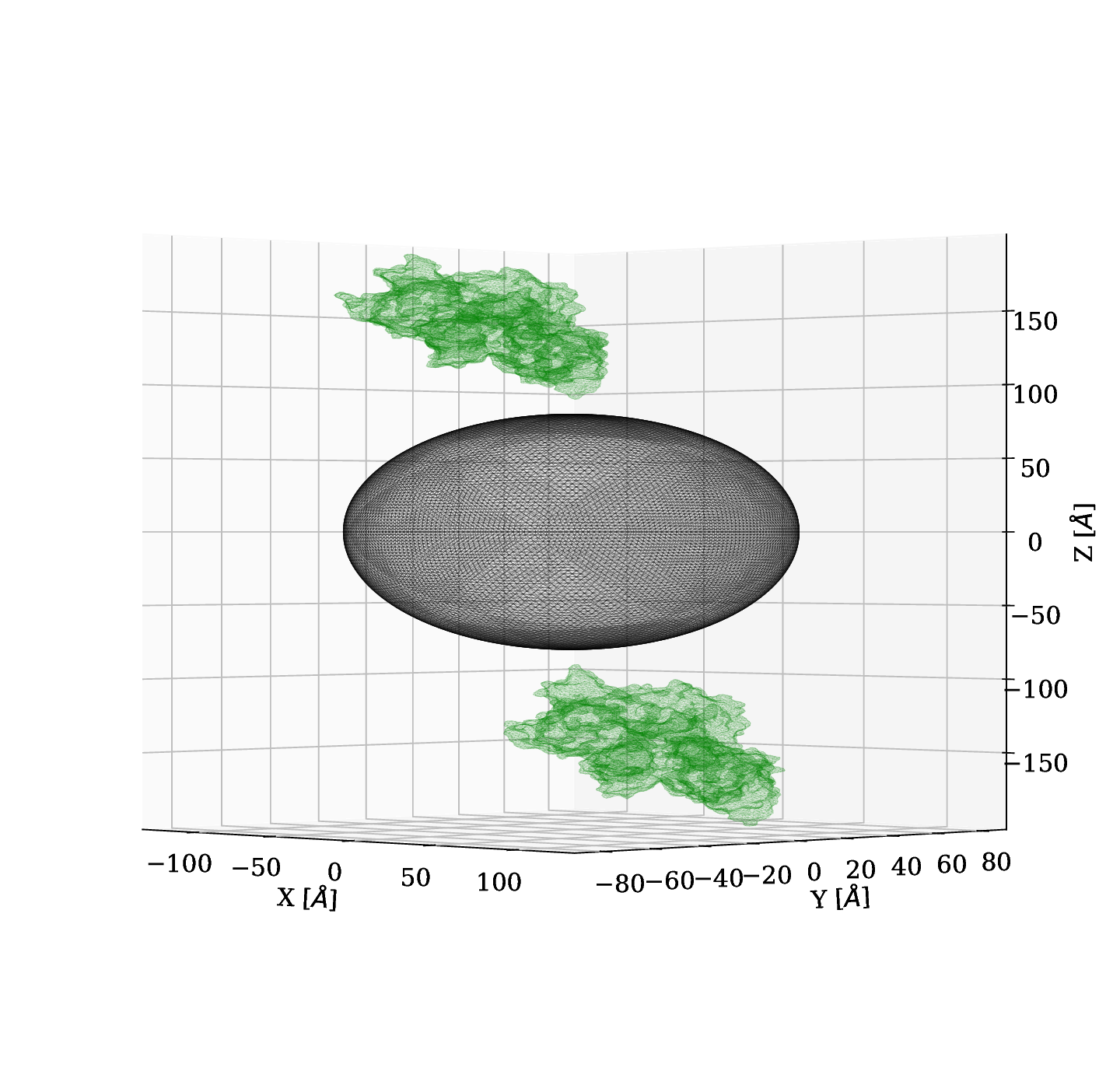

#### Fig 7. Close-up of spectrum shift d = 1 nm with protein tilted 45 deg.
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3901$ and it occurs at a wavelength of 384.5 nm
Shift: 0.5 nm
Results of 2 BSA located at 1 nm of +/- z with a rotation angle of 45 deg¶

#### Fig 8. Close-up of spectrum shift d = 1 nm with protein rotated 45 deg.
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3938$ and it occurs at a wavelength of 384.25 nm
Shift: 0.25 nm
Results of 2 BSA located at 1 nm of +/- z with a rotation angle of 90 deg¶

#### Fig 9. Close-up of spectrum shift d = 1 nm with protein rotated 90 deg.
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3943$ and it occurs at a wavelength of 384.25 nm
Shift: 0.25 nm
Results of 2 BSA located at 1 nm of +/- x¶

#### Fig 10. Close-up of spectrum shift d = 1 nm with proteins located at +/- x
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3979$ and it occurs at a wavelength of 384.0 nm
Shift: 0 nm
Results of 2 BSA located at 1 nm of +/- y¶

#### Fig 11. Close-up of spectrum shift d = 1 nm with proteins located at +/- y
Cext max at $d=\infty$ is $\sim 4010$ and it occurs at a wavelength of 384.0 nm.
Cext max at $d=1$ nm is $\sim 3985$ and it occurs at a wavelength of 384.0 nm
Shift: 0 nm
References¶
(1) Phan, Anh D., et al. "Surface plasmon resonances of protein-conjugated gold nanoparticles on graphitic substrates." Applied Physics Letters 103.16 (2013): 163702.
(2) Tang, Bin, et al. "Two-dimensional correlation localized surface plasmon resonance spectroscopy for analysis of the interaction between metal nanoparticles and bovine serum albumin." The Journal of Physical Chemistry C 114.49 (2010): 20990-20996.
#Ignore this cell, It simply loads a style for the notebook.
from IPython.core.display import HTML
def css_styling():
try:
styles = open("styles/custom.css", "r").read()
return HTML(styles)
except:
pass
css_styling()