using AIBECS
using PyPlot, PyCall
using LinearAlgebra
using GR, Interact
AIBECS.jl
The ideal tool for exploring global marine biogeochemical cycles
Algebraic Implicit Biogeochemical Elemental Cycling System
Check it on GitHub (look for AIBECS.jl)


with François Primeau and J. Keith Moore
Outline¶
- Motivation and concept
- Example 1: Radiocarbon
- Toy model circulation
- OCIM1
- Example 2: Phosphorus cycle
- AIBECS ecosystem
Motivation: Starting from the AWESOME OCIM¶
The AWESOME OCIM (for A Working Environment for Simulating Ocean Movement and Elemental cycling in an Ocean Circulation Inverse Model framework) by Seth John (USC)
Features: GUI, simple to use, fast, and good circulation
(thanks to the OCIM1 by Tim DeVries (UCSB))
Motivation: comes the AIBECS¶

Features (at present)
- simple to use
- fast
- Julia instead of MATLAB (free, open-source, better performance, and better syntax)
- nonlinear systems
- multiple tracers
- Other circulations (not just the OCIM)
- Parameter estimation/optimization and Sensitivity analysis (shameless plug: F-1 algorithm seminar tomorrow at the School of Mathematics)

AIBECS Concept: a simple interface¶
To build a BGC model with the AIBECS, you just need to
1. Specify the local sources and sinks
2. Chose the ocean circulation
(3. Specify the particle transport, if any)

AIBECS concept: Vectorization¶
The 3D ocean grid is rearranged
into a 1D column vector.
And linear operators are represented by matrices.
Example 1: Radiocarbon, a tracer for water age

Image credit: Luke Skinner, University of Cambridge
Tracer equation: transport + sources and sinks
The Tracer equation ($x=$ Radiocarbon concentration)
$$\frac{\partial x}{\partial t} + \color{RoyalBlue}{\nabla \cdot \left[ \boldsymbol{u} - \mathbf{K} \cdot \nabla \right]} x = \color{ForestGreen}{\underbrace{\Lambda(x)}_{\textrm{air–sea exchange}} - \underbrace{x / \tau}_{\textrm{radioactive decay}}}$$becomes
$$\frac{\partial \boldsymbol{x}}{\partial t} + \color{RoyalBlue}{\mathbf{T}} \, \boldsymbol{x} = \color{ForestGreen}{\mathbf{\Lambda}(\boldsymbol{x}) - \boldsymbol{x} / \tau}.$$with the transport matrix
Translating to AIBECS Code is easy¶
To use AIBECS, we must recast each tracer equation,
$$\frac{\partial \boldsymbol{x}}{\partial t} + \color{RoyalBlue}{\mathbf{T}} \, \boldsymbol{x} = \color{ForestGreen}{\mathbf{\Lambda}(\boldsymbol{x}) - \boldsymbol{x} / \tau}$$here, into the generic form:
$$\frac{\partial \boldsymbol{x}}{\partial t} + \color{RoyalBlue}{\mathbf{T}(\boldsymbol{p})} \, \boldsymbol{x} = \color{ForestGreen}{\boldsymbol{G}(\boldsymbol{x}, \boldsymbol{p})}$$where $\boldsymbol{p} =$ vector of model parameters
Circulation 1: The 2×2×2 Primeau model¶
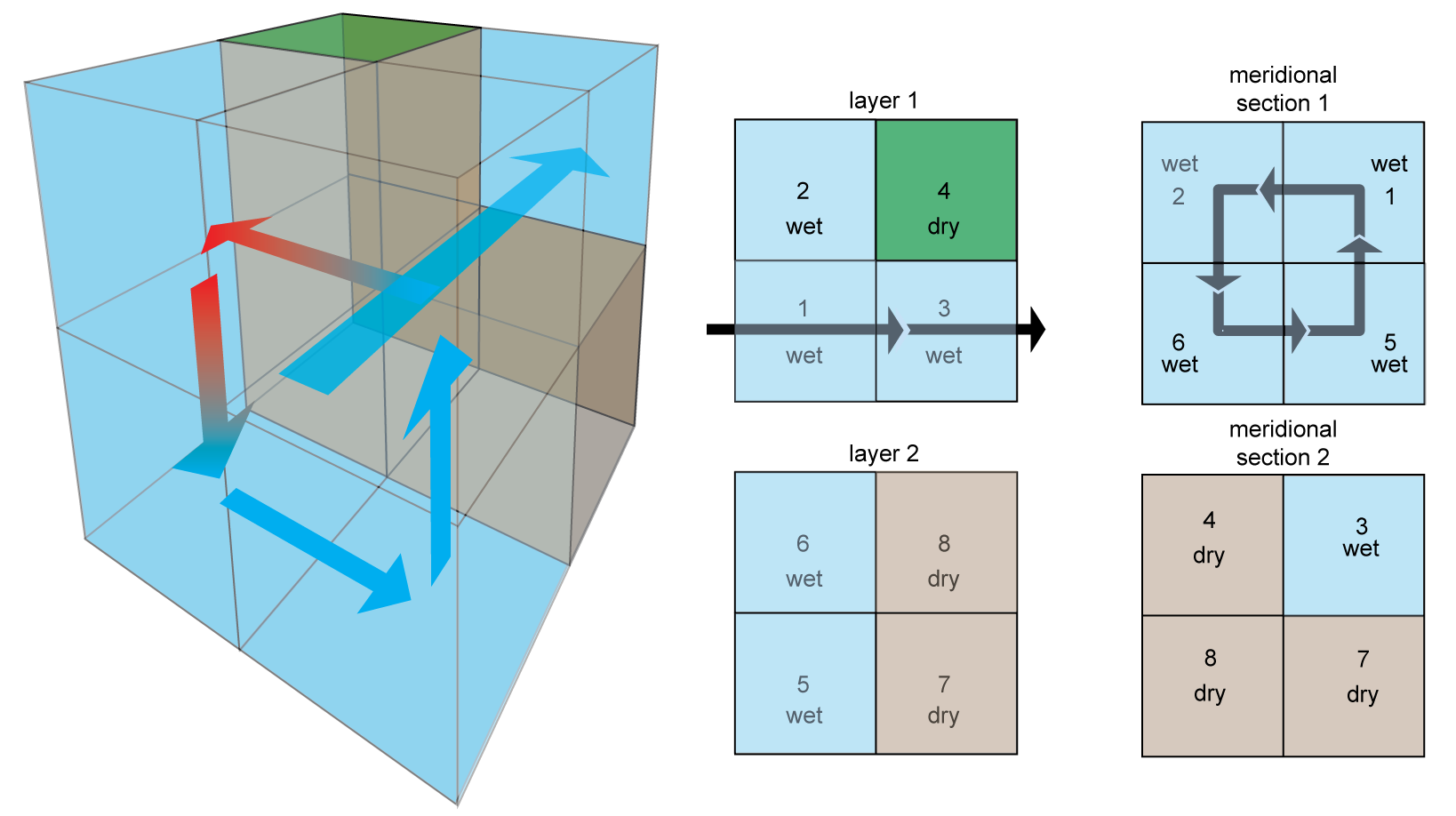
- ACC: Antarctic Circumpolar Current
- MOC: Meridional Overturning Circulation
- High-latitude mixing
(Credit: François Primeau, and Louis Primeau for the image)
Load the circulation via load:
wet3D, grd, T = Primeau_2x2x2.load();

wet3D is the mask of "wet" points
wet3D
grd is the grid of the circulation
grd
 We can check the depth of the boxes arranged in 3D
We can check the depth of the boxes arranged in 3D
grd.depth_3D
T
A sparse matrix is indexed by its non-zero values,
but we can check it out in full using Matrix:
Matrix(T)
Sources and sinks¶
Tracer equation reminder:
$$\frac{\partial \boldsymbol{x}}{\partial t} + \mathbf{T}(\boldsymbol{p}) \, \boldsymbol{x} = \boldsymbol{G}(\boldsymbol{x}, \boldsymbol{p})$$Let's write $\boldsymbol{G}(\boldsymbol{x}, \boldsymbol{p}) = \mathbf{\Lambda}(\boldsymbol{x}) - \boldsymbol{x} / \tau$
G(x,p) = Λ(x,p) - x / p.τ
Air–sea gas exchange¶
And define the air–sea gas exchange $\mathbf{\Lambda}(\boldsymbol{x}) = \frac{\lambda}{h} (R_\mathsf{atm} - \boldsymbol{x})$ at the surface with a piston velocity $\lambda$ over the top layer of height $h$
function Λ(x,p)
λ, h, Ratm = p.λ, p.h, p.Ratm
return @. λ / h * (Ratm - x) * (z == z₀)
end

Define z the depths in vector form.
(iwet converts from 3D to 1D)
iwet = findall(wet3D)
z = grd.depth_3D[iwet]
Define z₀ the depth of the top layer
z₀ = z[1]
So that z .== z₀ is true at the surface layer
z .== z₀
Model parameters¶
First, create a table of parameters using the AIBECS API
t = empty_parameter_table()
add_parameter!(t, :τ, 5730u"yr" / log(2)) # radioactive decay e-folding timescale
add_parameter!(t, :λ, 50u"m" / 10u"yr") # piston velocity
add_parameter!(t, :h, grd.δdepth[1]) # top layer height
add_parameter!(t, :Ratm, 1.0u"mol/m^3") # atmospheric concentration
t
Then, chose a name for the parameters (here C14_parameters), and create the vector p:
initialize_Parameters_type(t, "C14_parameters")
p = C14_parameters()
Note p has units!
In AIBECS, you give your parameters units and they are automatically converted to SI units under the hood.
(And they are converted back for pretty printing!)
State function (and Jacobian)¶
$$\frac{\partial \boldsymbol{x}}{\partial t} = \boldsymbol{G}(\boldsymbol{x}, \boldsymbol{p}) - \mathbf{T}(\boldsymbol{p}) \, \boldsymbol{x} = \color{Brown}{\boldsymbol{F}(\boldsymbol{x}, \boldsymbol{p})}$$We generate F and ∇ₓF via
F, ∇ₓF = state_function_and_Jacobian(p -> T, G) ;
The state function F(x,p)¶
Let's try F on a random state vector x
x = 0.5p.Ratm * ones(5)
F(x,p)
The Jacobian ∇ₓF¶
The Jacobian matrix is $\nabla_{\boldsymbol{x}}\boldsymbol{F}(\boldsymbol{x},\boldsymbol{p}) = \left[\frac{\partial F_i}{\partial x_j}\right]_{i,j}$, is useful for
- implicit time-steps
- solving the steady-state system
- optimization / uncertainty analysis
With AIBECS, the Jacobian is computed automatically for you under the hood... using dual numbers!
(Come to my Applied seminar tomorrow for more on dual numbers and... hyperdual numbers!)
Let's try ∇ₓF at x:
Matrix(∇ₓF(x,p))
Time stepping¶
AIBECS provides schemes for time-stepping
- Euler forward
- Euler backward
- Crank-Nicolson
- Crank-Nicolson leap-frog
Let's track the evolution of x through time
Define a function to apply the time steps n times for a time span of Δt starting from x₀
function time_steps(x₀, Δt, n, F, ∇ₓF)
x_hist = [x₀]
δt = Δt / n
for i in 1:n
push!(x_hist, AIBECS.crank_nicolson_step(last(x_hist), p, δt, F, ∇ₓF))
end
return reduce(hcat, x_hist), 0:δt:Δt
end
Let's run the model for 5000 years starting with x = 1 everywhere:
Δt = 5000u"yr" |> u"s" |> ustrip
x₀ = p.Ratm * ones(5)
x_hist, t_hist = time_steps(x₀, Δt, 1000, F, ∇ₓF)
Plotting the output is easy¶
The radiocarbon age, C14age, is given by $\log(R_{\mathrm{atm}}/\boldsymbol{x}) \tau$ because $\boldsymbol{x}\sim R_{\mathrm{atm}} \exp(-t/\tau)$
Let's plot its evolution with time:
C14age_hist = log.(p.Ratm ./ x_hist) * (p.τ * u"s" |> u"yr" |> ustrip)
PyPlot.figure(figsize=(8,4))
PyPlot.plot(t_hist .* 1u"s" .|> u"yr" .|> ustrip, C14age_hist')
PyPlot.xlabel("simulation time (years)")
PyPlot.ylabel("¹⁴C age (years)")
PyPlot.legend("box " .* string.(findall(vec(wet3D))))
PyPlot.title("Simulation of the evolution of ¹⁴C age with Crank-Nicolson time steps")
Solve directly for the steady state¶
Instead, we can directly solve for the steady-state, $\boldsymbol{s}$,
(using CTKAlg(), a quasi-Newton root-finding algorithm from C. T. Kelley)
i.e., such that $\boldsymbol{F}(\boldsymbol{s},\boldsymbol{p}) = 0$:
prob = SteadyStateProblem(F, ∇ₓF, x, p)
s = solve(prob, CTKAlg()).u
gives the age
log.(p.Ratm ./ s) * (p.τ * u"s" |> u"yr")
35'000 years without the steady-state solver!¶
How long would it take to reach that steady-state with time-stepping?
We chan check by tracking the norm of F(x,p):
Δt = 40000u"yr" |> u"s" |> ustrip
x_hist, t_hist = time_steps(x₀, Δt, 4000, F, ∇ₓF)
PyPlot.figure(figsize=(7,3))
PyPlot.semilogy(t_hist .* 1u"s" .|> u"yr" .|> ustrip, [norm(F(s,p)) for i in 1:size(x_hist,2)], label="steady-state")
PyPlot.semilogy(t_hist .* 1u"s" .|> u"yr" .|> ustrip, [norm(F(x_hist[:,i],p)) for i in 1:size(x_hist,2)], label="time-stepping")
PyPlot.xlabel("simulation time (years)"); PyPlot.ylabel("|F(x,p)| (mol m⁻³ s⁻¹)");
PyPlot.legend(); PyPlot.title("Stability of ¹⁴C age with simulation time")
Circulation 2: OCIM1¶
The Ocean Circulation Inverse Model (OCIM) version 1 is loaded via
wet3D, grd, T = OCIM1.load() ;
Redefine F and ∇ₓF for the new T:
F, ∇ₓF = state_function_and_Jacobian(p -> T, G) ;
Redefine iwet and x for the new grid size
iwet = findall(wet3D)
x = p.Ratm * ones(length(iwet))
Redefine h, z₀, and z for the new grid
p.h = grd.δdepth[1] |> upreferred |> ustrip
z = grd.depth_3D[iwet]
z₀ = z[1]
Solve for the steady-state of radio carbon and convert to age
prob = SteadyStateProblem(F, ∇ₓF, x, p)
s = solve(prob, CTKAlg()).u
C14age = log.(p.Ratm ./ s) * p.τ * u"s" .|> u"yr"
And plot horizontal slices using GR and Interact:
lon, lat = grd.lon |> ustrip, grd.lat |> ustrip
function horizontal_slice(x, levels, title)
x_3D = fill(NaN, size(grd))
x_3D[iwet] .= x .|> ustrip
GR.figure(figsize=(10,5))
@manipulate for iz in 1:size(grd)[3]
GR.xlim([0,360])
GR.title(string(title, " at $(AIBECS.round(grd.depth[iz])) depth"))
GR.contourf(lon, lat, x_3D[:,:,iz]', levels=levels)
end
end
horizontal_slice(C14age, 0:100:2400, "14C age [yr] using the OCIM1 circulation")
Or zonal slices:
function zonal_slice(x, levels, title)
x_3D = fill(NaN, size(grd))
x_3D[iwet] .= x .|> ustrip
GR.figure(figsize=(10,5))
@manipulate for longitude in (grd.lon .|> ustrip)
GR.title(string(title, " at $(round(longitude))°"))
ilon = findfirst(ustrip.(grd.lon) .== longitude)
GR.contourf(lat, reverse(-grd.depth .|> ustrip), reverse(x_3D[:,ilon,:], dims=2), levels=levels)
end
end
zonal_slice(C14age, 0:100:2400, "14C age [yr] using the OCIM1 circulation")

Example 2: A phosphorus cycle
Dissolved inorganic phosphrous (DIP)
(transported by the ocean circulation)
and particulate organic phosphrous (POP)
(transported by sinking particles)
Ocean circulation¶
For DIP, the advective–eddy-diffusive transport operator, $\nabla \cdot (\boldsymbol{u} + \mathbf{K}\cdot\nabla)$, is converted into the matrix T:
T_DIP(p) = T
Sinking particles¶
For POP, the particle flux divergence (PFD) operator, $\nabla \cdot \boldsymbol{w}$, is created via buildPFD:
T_POP(p) = buildPFD(grd, wet3D, sinking_speed = w(p))
The settling velocity, w(p), is assumed linearly increasing with depth z to yield a "Martin curve profile"
w(p) = p.w₀ .+ p.w′ * (z .|> ustrip)
relu(x) = (x .≥ 0) .* x
zₑ = 80u"m" # depth of the euphotic zone
function uptake(DIP, p)
τ, k = p.τ, p.k
DIP⁺ = relu(DIP)
return 1/τ * DIP⁺.^2 ./ (DIP⁺ .+ k) .* (z .≤ zₑ)
end
remineralization(POP, p) = p.κ * POP
geores(x, p) = (p.xgeo .- x) / p.τgeo
G_DIP(DIP, POP, p) = -uptake(DIP, p) + remineralization(POP, p) + geores(DIP, p)
G_POP(DIP, POP, p) = uptake(DIP, p) - remineralization(POP, p)
Parameters¶
t = empty_parameter_table() # empty table of parameters
add_parameter!(t, :xgeo, 2.12u"mmol/m^3")
add_parameter!(t, :τgeo, 1.0u"Myr")
add_parameter!(t, :k, 6.62u"μmol/m^3")
add_parameter!(t, :w₀, 0.64u"m/d")
add_parameter!(t, :w′, 0.13u"1/d")
add_parameter!(t, :κ, 0.19u"1/d")
add_parameter!(t, :τ, 236.52u"d")
initialize_Parameters_type(t, "Pcycle_Parameters") # Generate the parameter type
p = Pcycle_Parameters()
State function F and Jacobian ∇ₓF¶
nb = length(iwet); x = ones(2nb)
F, ∇ₓF = state_function_and_Jacobian((T_DIP,T_POP), (G_DIP,G_POP), nb) ;
Solve the steady-state PDE system
prob = SteadyStateProblem(F, ∇ₓF, x, p)
s = solve(prob, CTKAlg(), preprint=" ").u
Plot DIP
DIP, POP = state_to_tracers(s, nb, 2)
DIP_unit = u"mmol/m^3"
horizontal_slice(DIP * u"mol/m^3" .|> DIP_unit, 0:0.3:3, "P-cycle model: DIP [mmol m^{-3}]")
Plot POP
zonal_slice(POP .|> relu .|> log10, -10:1:10, "P-cycle model: POP [log\\(mol m^{-3}\\)]")
We can also make publication-quality plots using Python's Cartopy
ccrs = pyimport("cartopy.crs")
cfeature = pyimport("cartopy.feature")
lon_cyc = [lon; 360+lon[1]]
DIP_3D = fill(NaN, size(grd)); DIP_3D[iwet] = DIP * u"mol/m^3" .|> u"mmol/m^3" .|> ustrip
DIP_3D_cyc = cat(DIP_3D, DIP_3D[:,[1],:], dims=2)
f1 = PyPlot.figure(figsize=(12,5))
@manipulate for iz in 1:24
withfig(f1, clear=true) do
ax = PyPlot.subplot(projection = ccrs.EqualEarth(central_longitude=-155.0))
ax.add_feature(cfeature.COASTLINE, edgecolor="#000000") # black coast lines
ax.add_feature(cfeature.LAND, facecolor="#CCCCCC") # gray land
plt = PyPlot.contourf(lon_cyc, lat, DIP_3D_cyc[:,:,iz], levels=0:0.1:3.5,
transform=ccrs.PlateCarree(), zorder=-1, extend="both")
plt2 = PyPlot.contour(plt, lon_cyc, lat, DIP_3D_cyc[:,:,iz], levels=plt.levels[1:5:end],
transform=ccrs.PlateCarree(), zorder=0, extend="both", colors="k")
ax.clabel(plt2, fmt="%2.1f", colors="k", fontsize=14)
cbar = PyPlot.colorbar(plt, orientation="vertical", extend="both", ticks=plt2.levels)
cbar.add_lines(plt2)
cbar.ax.tick_params(labelsize=14)
cbar.set_label(label="mmol / m³", fontsize=16)
PyPlot.title("Publication-quality DIP with Cartopy; depth = $(string(round(typeof(1u"m"),grd.depth[iz])))", fontsize=20)
end
end

