Wiggle nuclei¶
In this notebook we move the positions of nuclei artifically to afterwards segment them again and draw a mesh between them. The mesh will change more or less depending on how much we modify the nuclei positions randomly.
In [1]:
# !pip install napari-animated-gif-io pyclesperanto-prototype stackview
In [2]:
import numpy as np
import pyclesperanto_prototype as cle
from napari_animated_gif_io._function import save_as_animated_gif
cle.select_device("tx")
Out[2]:
<NVIDIA GeForce RTX 3050 Ti Laptop GPU on Platform: NVIDIA CUDA (1 refs)>
Load data¶
Tribolium castaneum, nuclei-GFP transgenic line, cropped. Image data source: Daniela Vorkel, Myers lab, MPI-CBG / CSBD
https://github.com/clEsperanto/clesperanto_example_data/blob/main/Lund_000500_resampled-cropped.tif
In [3]:
image = cle.imread("../../../clesperanto_example_data/Lund_000500_resampled-cropped.tif")
image
Out[3]:
|
|
cle._ image
|
Segment nuclei¶
In [4]:
nuclei = cle.voronoi_otsu_labeling(cle.top_hat_box(image, radius_x=5, radius_y=5), spot_sigma=1)
nuclei
Out[4]:
|
|
cle._ image
|
In [5]:
def move_stuff_around(image, nuclei, distance=1):
"""
Randomly move positions of given nuclei by applying a vector field to the image.
"""
centroids = cle.reduce_labels_to_centroids(nuclei)
num_nuclei = nuclei.max()
vector_field_x = make_random_vector_field(nuclei, distance)
vector_field_y = make_random_vector_field(nuclei, distance)
vector_field_z = make_random_vector_field(nuclei, distance)
return cle.apply_vector_field(image, vector_field_x, vector_field_y, vector_field_z)
def make_random_vector_field(nuclei, distance):
"""
Renerate a vector field from a label image by randomly introducing shift according to a given distance.
"""
random_shifts = (np.random.random((1, int(num_nuclei + 1))) * 2 -1) * distance
random_shifts[0,0] = 0 # keep background where it is
vector_field_sharp = cle.replace_intensities(nuclei, random_shifts)
sigma = 10
return cle.gaussian_blur(vector_field_sharp, sigma_x=sigma, sigma_y=sigma, sigma_z=sigma)
In [6]:
num_nuclei = nuclei.max()
num_nuclei
Out[6]:
285.0
In [7]:
def analyze(image, nuclei):
image_to_analyze = move_stuff_around(image, nuclei, distance=10)
specific_nuclei = cle.voronoi_otsu_labeling(cle.top_hat_box(image_to_analyze, radius_x=5, radius_y=5), spot_sigma=1)
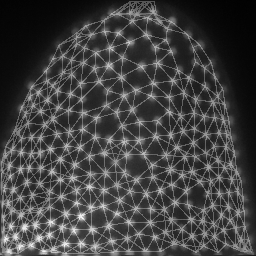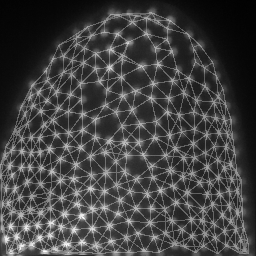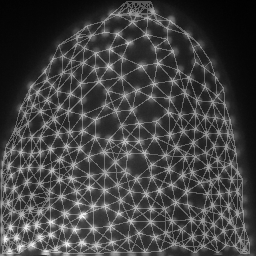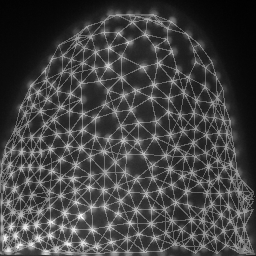
mesh = cle.draw_mesh_between_proximal_labels(specific_nuclei, maximum_distance=25)
return mesh * image_to_analyze.max()/4 + image_to_analyze
In [8]:
images = np.asarray([cle.maximum_z_projection(analyze(image, nuclei)) for i in range(10)])
In [9]:
save_as_animated_gif(images, "visualization.gif")
Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 812.25]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 805.25]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 813.0]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 812.25]. Convert image to uint8 prior to saving to suppress this warning. Lossy conversion from float32 to uint8. Range [141.0, 806.0]. Convert image to uint8 prior to saving to suppress this warning.
In [10]:
import stackview
stackview.slice(np.asarray(images), continuous_update=True)
Out[10]:
VBox(children=(HBox(children=(VBox(children=(ImageWidget(height=256, width=256),)),)), IntSlider(value=5, desc…