Clean Bibliography¶
To goal of this notebook is to clean your .bib file to ensure that it only contains the full first names of references that you have cited in your paper. The full first names will then be used to query the probabilistic gender classifier, Gender API. The full names will be used to query for probabilistic race using the ethnicolr package.
The only required file you need is your manuscript's bibliography in .bib format. Your .bib must only contain references cited in the manuscript. Otherwise, the estimated proportions will be inaccurate.
If you intend to analyze the reference list of a published paper instead of your own manuscript in progress, search the paper on Web of Knowledge (you will need institutional access). Next, download the .bib file from Web of Science following these instructions, but start from Step 4 and on Step 6 select BibTeX instead of Plain Text.
If you are not using LaTeX, collect and organize only the references you have cited in your manuscript using your reference manager of choice (e.g. Mendeley, Zotero, EndNote, ReadCube, etc.) and export that selected bibliography as a .bib file. Please try to export your .bib in an output style that uses full first names (rather than only first initials) and using the full author lists (rather than abbreviated author lists with "et al."). If first initials are included, our code will automatically retrieve about 70% of those names using the article title or DOI.
- Export
.bibfrom Mendeley - Export
.bibfrom Zotero - Export
.bibfrom EndNote. Note: Please export full first names by either choosing an output style that does so by default (e.g. in MLA style) or by customizing an output style. - Export
.bibfrom Read Cube Papers
For those working in LaTeX, we can use an optional .aux file to automatically filter your .bib to check that it only contains entries which are cited in your manuscript.
| Input | Output |
|---|---|
.bib file(s)(REQUIRED) |
cleanBib.csv: table of author first names, titles, and .bib keys |
.aux file (OPTIONAL) |
predictions.csv: table of author first names, estimated gender classification, and confidence |
.tex file (OPTIONAL) |
race_gender_citations.pdf: heat map of your citations broken down by probabilistic gender and race estimations |
yourTexFile_gendercolor.tex: your .tex file modified to compile .pdf with in-line citations colored-coded by gender pairs |
1. Import functions¶
Upload your .bib file(s) and optionally an .aux file generated from compiling your LaTeX manuscript and your .tex file
Then, run the code block below. (click to select the block and then press Ctrl+Enter; or click the block and press the Run button in the top menubar)
import numpy as np
import bibtexparser
from bibtexparser.bparser import BibTexParser
import glob
import subprocess
import os
from pybtex.database.input import bibtex
import csv
from pylatexenc.latex2text import LatexNodes2Text
import unicodedata
import re
import pandas as pd
from habanero import Crossref
import string
from time import sleep
import tqdm
import matplotlib.pylab as plt
import matplotlib.gridspec as gridspec
import json
import pickle
from urllib.request import urlopen
from ethnicolr import census_ln, pred_census_ln,pred_wiki_name
from pybtex.database import parse_file
import seaborn as sns
def checkcites_output(aux_file):
'''take in aux file for tex document, return list of citation keys
that are in .bib file but not in document'''
result = subprocess.run(['texlua', 'checkcites.lua', aux_file[0]], stdout=subprocess.PIPE)
result = result.stdout.decode('utf-8')
unused_array_raw = result.split('\n')
# process array of unused references + other output
unused_array_final = list()
for x in unused_array_raw:
if len(x) > 0: # if line is not empty
if x[0] == '-': # and if first character is a '-', it's a citation key
unused_array_final.append(x[2:]) # truncate '- '
if "------------------------------------------------------------------------" in unused_array_final:
return(result)
else:
return(unused_array_final)
def removeMiddleName(line):
arr = line.split()
last = arr.pop()
n = len(arr)
if n == 4:
first, middle = ' '.join(arr[:2]), ' '.join(arr[2:])
elif n == 3:
first, middle = arr[0], ' '.join(arr[1:])
elif n == 2:
first, middle = arr
elif n==1:
return line
return(str(first + ' ' + middle))
def returnFirstName(line):
arr = line.split()
n = len(arr)
if n == 4:
first, middle = ' '.join(arr[:2]), ' '.join(arr[2:])
elif n == 3:
first, middle = arr[0], ' '.join(arr[1:])
elif n == 2:
first, middle = arr
elif n==1:
return line
return(str(middle))
def convertLatexSpecialChars(latex_text):
return LatexNodes2Text().latex_to_text(latex_text)
def convertSpecialCharsToUTF8(text):
data = LatexNodes2Text().latex_to_text(text)
return unicodedata.normalize('NFD', data).encode('ascii', 'ignore').decode('utf-8')
def namesFromXref(doi, title, authorPos):
'''Use DOI and article titles to query Crossref for author list'''
if authorPos == 'first':
idx = 0
elif authorPos == 'last':
idx = -1
# get cross ref data
authors = ['']
# first try DOI
if doi != "":
works = cr.works(query = title, select = ["DOI","author"], limit=1, filter = {'doi': doi})
if works['message']['total-results'] > 0:
authors = works['message']['items'][0]['author']
elif title != '':
works = cr.works(query = f'title:"{title}"', select = ["title","author"], limit=10)
cnt = 0
name = ''
# check that you grabbed the proper paper
if works['message']['items'][cnt]['title'][0].lower() == title.lower():
authors = works['message']['items'][0]['author']
# check the all fields are available
if not 'given' in authors[idx]:
name = ''
else:
# trim initials
name = authors[idx]['given'].replace('.',' ').split()[0]
return name
def namesFromXrefSelfCite(doi, title):
selfCiteCheck = 0
# get cross ref data
authors = ['']
# first try DOI
if doi != "":
works = cr.works(query = title, select = ["DOI","author"], limit=1, filter = {'doi': doi})
if works['message']['total-results'] > 0:
authors = works['message']['items'][0]['author']
for i in authors:
if i != "":
first = i['given'].replace('.',' ').split()[0]
last = i['family'].replace('.',' ').split()[0]
authors = removeMiddleName(last + ", " + first)
if authors in removeMiddleName(yourFirstAuthor) or authors in removeMiddleName(convertSpecialCharsToUTF8(yourFirstAuthor)) or authors in removeMiddleName(yourLastAuthor) or authors in removeMiddleName(convertSpecialCharsToUTF8(yourLastAuthor)):
selfCiteCheck += 1
return selfCiteCheck
cr = Crossref()
homedir = '/home/jovyan/'
bib_files = glob.glob(homedir + '*.bib')
paper_aux_file = glob.glob(homedir + '*.aux')
paper_bib_file = 'library_paper.bib'
try:
tex_file = glob.glob(homedir + "*.tex")[0]
except:
print('No optional .tex file found.')
2. Define the first and last author of your paper.¶
For example:
yourFirstAuthor = 'Teich, Erin G.'
yourLastAuthor = 'Bassett, Danielle S.'
And optionally, define any co-first or co-last author(s), making sure to keep the square brackets to define a list.
For example:
optionalEqualContributors = ['Dworkin, Jordan', 'Stiso, Jennifer']
or
optionalEqualContributors = ['Dworkin, Jordan']
If you are analyzing published papers' reference lists from Web of Science, change the variable checkingPublishedArticle to True:
checkingPublishedArticle = True
Then, run the code block below. (click to select the block and then press Ctrl+Enter; or click the block and press the Run button in the top menubar)
yourFirstAuthor = 'LastName, FirstName OptionalMiddleInitial'
yourLastAuthor = 'LastName, FirstName OptionalMiddleInitial'
optionalEqualContributors = ['LastName, FirstName OptionalMiddleInitial', 'LastName, FirstName OptionalMiddleInitial']
checkingPublishedArticle = False
if (yourFirstAuthor == 'LastName, FirstName OptionalMiddleInitial') or (yourLastAuthor == 'LastName, FirstName OptionalMiddleInitial'):
raise ValueError("Please enter your manuscript's first and last author names")
if paper_aux_file:
if optionalEqualContributors == ['LastName, FirstName OptionalMiddleInitial', 'LastName, FirstName OptionalMiddleInitial']:
citing_authors = np.array([yourFirstAuthor, yourLastAuthor])
else:
citing_authors = np.array([yourFirstAuthor, yourLastAuthor, optionalEqualContributors])
print(checkcites_output(paper_aux_file))
unused_in_paper = checkcites_output(paper_aux_file) # get citations in library not used in paper
print("Unused citations: ", unused_in_paper.count('=>'))
parser = BibTexParser()
parser.ignore_nonstandard_types = False
parser.common_strings = True
bib_data = None
for bib_file in bib_files:
with open(bib_file) as bibtex_file:
if bib_data is None:
bib_data = bibtexparser.bparser.BibTexParser(common_strings=True, ignore_nonstandard_types=False).parse_file(bibtex_file)
else:
bib_data_extra = bibtexparser.bparser.BibTexParser(common_strings=True, ignore_nonstandard_types=False).parse_file(bibtex_file)
bib_data.entries_dict.update(bib_data_extra.entries_dict)
bib_data.entries.extend(bib_data_extra.entries)
all_library_citations = list(bib_data.entries_dict.keys())
print("All citations: ", len(all_library_citations))
for k in all_library_citations:
if re.search('\\b'+ k + '\\b', unused_in_paper.replace('\n',' ').replace('=>',' ')) != None:
del bib_data.entries_dict[k] # remove from entries dictionary if not in paper
in_paper_mask = [re.search('\\b'+ bib_data.entries[x]['ID'] + '\\b', unused_in_paper.replace('\n',' ').replace('=>',' ')) == None for x in range(len(bib_data.entries))]
bib_data.entries = [bib_data.entries[x] for x in np.where(in_paper_mask)[0]] # replace entries list with entries only in paper
del bib_data.comments
duplicates = []
for key in bib_data.entries_dict.keys():
count = str(bib_data.entries).count("'ID\': \'"+ key + "\'")
if count > 1:
duplicates.append(key)
if len(duplicates) > 0:
raise ValueError("In your .bib file, please remove duplicate entries or duplicate entry ID keys for:", ' '.join(map(str, duplicates)))
if os.path.exists(paper_bib_file):
os.remove(paper_bib_file)
with open(paper_bib_file, 'w') as bibtex_file:
bibtexparser.dump(bib_data, bibtex_file)
# define first author and last author names of citing paper -- will exclude citations of these authors
# beware of latex symbols within author names
# in_paper_citations = list(bib_data.entries_dict.keys())
in_paper_citations = [bib_data.entries[x]['ID'] for x in range(len(bib_data.entries))] # get list of citation keys in paper
# extract author list for every cited paper
cited_authors = [bib_data.entries_dict[x]['author'] for x in in_paper_citations]
# find citing authors in cited author list
# using nested list comprehension, make a citing author -by- citation array of inclusion
self_cite_mask = np.array([[str(citing_author) in authors for authors in cited_authors] for citing_author in citing_authors])
self_cite_mask = np.any(self_cite_mask,axis=0) # collapse across citing authors such that any coauthorship by either citing author -> exclusion
print("Self-citations: ", [bib_data.entries[x]['ID'] for x in np.where(self_cite_mask)[0]]) # print self citations
for idx,k in enumerate(in_paper_citations):
if self_cite_mask[idx]:
del bib_data.entries_dict[k] # delete citation from dictionary if self citationi
bib_data.entries = [bib_data.entries[x] for x in np.where(np.invert(self_cite_mask))[0]] # replace entries list with entries that aren't self citations
paper_bib_file_excl_sc = os.path.splitext(paper_bib_file)[0] + '_noselfcite.bib'
if os.path.exists(paper_bib_file_excl_sc):
os.remove(paper_bib_file_excl_sc)
with open(paper_bib_file_excl_sc, 'w') as bibtex_file:
bibtexparser.dump(bib_data, bibtex_file)
if os.path.exists('*_noselfcite.bib'):
ID = glob.glob(homedir + paper_bib_file_excl_sc)
else:
ID = glob.glob(homedir + '*bib')
with open(ID[0]) as bibtex_file:
bib_data = bibtexparser.bparser.BibTexParser(common_strings=True, ignore_nonstandard_types=False).parse_file(bibtex_file)
duplicates = []
for key in bib_data.entries_dict.keys():
count = str(bib_data.entries).count("'ID\': \'"+ key + "\'")
if count > 1:
duplicates.append(key)
if len(duplicates) > 0:
raise ValueError("In your .bib file, please remove duplicate entries or duplicate entry ID keys for:", ' '.join(map(str, duplicates)))
if checkingPublishedArticle == True:
FA = []
LA = []
counter = 1
selfCiteCount = 0
titleCount = 1 #
counterNoDOI = list() # row index (titleCount) of entries with no DOI
outPath = homedir + 'cleanedBib.csv'
if os.path.exists(outPath):
os.remove(outPath)
with open(outPath, 'w', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',', quotechar='|', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['Article', 'FA', 'LA', 'Title', 'SelfCite', 'CitationKey'])
citedArticleDOI = list()
citedArticleNoDOI = list()
allArticles = list()
for entry in bib_data.entries:
my_string= entry['cited-references'].split('\n')
for citedArticle in my_string:
allArticles.append(citedArticle)
if citedArticle.partition("DOI ")[-1]=='':
citedArticleNoDOI.append(citedArticle)
counterNoDOI.append(titleCount)
else:
line = citedArticle.partition("DOI ")[-1].replace("DOI ","").rstrip(".")
line = ''.join( c for c in line if c not in '{[}] ')
if "," in line:
line = line.partition(",")[-1]
citedArticleDOI.append(line)
with open('citedArticlesDOI.csv', 'a', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow([line])
titleCount += 1
articleNum = 0
for doi in citedArticleDOI:
try:
FA = namesFromXref(doi, '', 'first')
except UnboundLocalError:
sleep(1)
continue
try:
LA = namesFromXref(doi, '', 'last')
except UnboundLocalError:
sleep(1)
continue
try:
selfCiteCount = namesFromXrefSelfCite(doi, '')
except UnboundLocalError:
sleep(1)
continue
with open(outPath, 'a', newline='') as csvfile:
if selfCiteCount == 0:
writer = csv.writer(csvfile, delimiter=',')
getArticleIndex = [i for i, s in enumerate(allArticles) if doi in s]
writer.writerow([counter, convertSpecialCharsToUTF8(FA), convertSpecialCharsToUTF8(LA), allArticles[[i for i, s in enumerate(allArticles) if doi in s][0]], '', ''])
print(str(counter) + ": " + doi )
counter += 1
else:
print(str(articleNum) + ": " + doi + "\t\t\t <-- self-citation" )
articleNum += 1
if len(citedArticleNoDOI)>0:
print()
for elem in citedArticleNoDOI:
with open(outPath, 'a', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow([counter, '', '', elem, '', ''])
print(str(counter) + ": " + elem )
counter += 1
print()
raise ValueError("WARNING: No article DOI was provided for the last " + str(len(citedArticleNoDOI)) + " listed papers. Please manually search for these articles. IF AND ONLY IF your citing paper's first and last author are not co-authors in the paper that was cited, enter the first name of the first and last authors of the paper that was cited manually. Then, continue to the next code block.")
else:
FA = []
LA = []
parser = bibtex.Parser()
bib_data = parser.parse_file(ID[0])
counter = 1
nameCount = 0
outPath = homedir + 'cleanedBib.csv'
if os.path.exists(outPath):
os.remove(outPath)
with open(outPath, 'w', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',', quotechar='|', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['Article', 'FA', 'LA', 'Title', 'SelfCite', 'CitationKey'])
for key in bib_data.entries.keys():
diversity_bib_titles = ['The extent and drivers of gender imbalance in neuroscience reference lists','The gender citation gap in international relations','Quantitative evaluation of gender bias in astronomical publications from citation counts', '\# CommunicationSoWhite', '{Just Ideas? The Status and Future of Publication Ethics in Philosophy: A White Paper}','Gendered citation patterns across political science and social science methodology fields','Gender Diversity Statement and Code Notebook v1.0']
if bib_data.entries[key].fields['title'] in diversity_bib_titles:
continue
try:
author = bib_data.entries[key].persons['author']
except:
author = bib_data.entries[key].persons['editor']
FA = author[0].rich_first_names
LA = author[-1].rich_first_names
FA = convertLatexSpecialChars(str(FA)[7:-3]).translate(str.maketrans('', '', string.punctuation)).replace('Protected',"").replace(" ",'')
LA = convertLatexSpecialChars(str(LA)[7:-3]).translate(str.maketrans('', '', string.punctuation)).replace('Protected',"").replace(" ",'')
# check that we got a name (not an initial) from the bib file, if not try using the title in the crossref API
try:
title = bib_data.entries[key].fields['title'].replace(',', '').replace(',', '').replace('{','').replace('}','')
except:
title = ''
try:
doi = bib_data.entries[key].fields['doi']
except:
doi = ''
if FA == '' or len(FA.split('.')[0]) <= 1:
while True:
try:
FA = namesFromXref(doi, title, 'first')
except UnboundLocalError:
sleep(1)
continue
break
if LA == '' or len(LA.split('.')[0]) <= 1:
while True:
try:
LA = namesFromXref(doi, title, 'last')
except UnboundLocalError:
sleep(1)
continue
break
if (yourFirstAuthor!='LastName, FirstName OptionalMiddleInitial') and (yourLastAuthor!='LastName, FirstName OptionalMiddleInitial'):
selfCiteCheck1 = [s for s in author if removeMiddleName(yourLastAuthor) in str([convertLatexSpecialChars(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertLatexSpecialChars(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
selfCiteCheck1a = [s for s in author if removeMiddleName(yourLastAuthor) in str([convertSpecialCharsToUTF8(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertSpecialCharsToUTF8(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
selfCiteCheck1b = [s for s in author if removeMiddleName(yourLastAuthor) in str([convertSpecialCharsToUTF8(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), LA]).replace("'", "")]
selfCiteCheck2 = [s for s in author if removeMiddleName(yourFirstAuthor) in str([convertLatexSpecialChars(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertLatexSpecialChars(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
selfCiteCheck2a = [s for s in author if removeMiddleName(yourFirstAuthor) in str([convertSpecialCharsToUTF8(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertSpecialCharsToUTF8(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
selfCiteCheck2b = [s for s in author if removeMiddleName(yourFirstAuthor) in str([convertSpecialCharsToUTF8(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), FA]).replace("'", "")]
nameCount = 0
if optionalEqualContributors != ('LastName, FirstName OptionalMiddleInitial', 'LastName, FirstName OptionalMiddleInitial'):
for name in optionalEqualContributors:
selfCiteCheck3 = [s for s in author if removeMiddleName(name) in str([convertLatexSpecialChars(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertLatexSpecialChars(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
selfCiteCheck3a = [s for s in author if removeMiddleName(name) in str([convertSpecialCharsToUTF8(str(s.rich_last_names)[7:-3]).replace("', Protected('","").replace("'), '", ""), convertSpecialCharsToUTF8(str(s.rich_first_names)[7:-3]).replace("', Protected('","").replace("'), '", "")]).replace("'", "")]
if len(selfCiteCheck3)>0:
nameCount += 1
if len(selfCiteCheck3a)>0:
nameCount += 1
selfCiteChecks = [selfCiteCheck1, selfCiteCheck1a, selfCiteCheck1b, selfCiteCheck2, selfCiteCheck2a, selfCiteCheck2b]
if sum([len(check) for check in selfCiteChecks]) + nameCount > 0:
selfCite = 'Y'
if len(FA) < 2:
print(str(counter) + ": " + key + "\t\t <-- self-citation <-- ***NAME MISSING OR POSSIBLY INCOMPLETE***")
else:
print(str(counter) + ": " + key + " <-- self-citation")
else:
selfCite= 'N'
if len(FA) < 2:
print(str(counter) + ": " + key + "\t\t <-- ***NAME MISSING OR POSSIBLY INCOMPLETE***")
else:
print(str(counter) + ": " + key)
else:
selfCite = 'NA'
with open(outPath, 'a', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',', quotechar='|', quoting=csv.QUOTE_MINIMAL)
writer.writerow([counter, convertSpecialCharsToUTF8(FA), convertSpecialCharsToUTF8(LA), title, selfCite, key])
counter += 1
3. Estimate gender and race of authors from cleaned bibliography¶
Checkpoint for cleaned bibliography and using Gender API to estimate genders by first names¶
After registering for a gender-api (free account available), use your 500 free monthly search credits by pasting your API key in the code for the line indicated below (replace only YOUR ACCOUNT KEY HERE):
genderAPI_key <- '&key=YOUR ACCOUNT KEY HERE'
You can find your key in your account's profile page.
NOTE: Please edit your .bib file using information printed by the code and provided in cleanedBib.csv. Edit directly within the Binder environment by clicking the Edit button, making modifications, and saving the file.
Common issues include:
- Bibliography entry did not include a last author because the author list was truncated by "and Others" or "et al."
- Some older journals articles only provide first initial and not full first names, in which case you will need to go digging via Google to identify that person.
- In rare cases where the author cannot be identified even after searching by hand, replace the first name with "UNKNOWNNAME" so that the classifier will estimate the gender as unknown.
NOTE: your free account has 500 queries per month. This box contains the code that will use your limited API credits/queries if it runs without error. Re-running all code repeatedly will repeatedly use these credits.
Then, run the code blocks below. (click to select the block and then press Ctrl+Enter; or click the block and press the Run button in the top menubar)
genderAPI_key <- '&key=YOUR ACCOUNT KEY HERE'
fileConn<-file("genderAPIkey.txt")
writeLines(c(genderAPI_key), fileConn)
close(fileConn)
names=read.csv("/home/jovyan/cleanedBib.csv",stringsAsFactors=F)
setwd('/home/jovyan/')
require(rjson)
gendFA=NULL;gendLA=NULL
gendFA_conf=NULL;gendLA_conf=NULL
namesIncompleteFA=NULL
namesIncompleteLA=NULL
incompleteKeys=list()
incompleteRows=list()
for(i in 1:nrow(names)){
if (nchar(names$FA[i])<2 || grepl("\\.", names$FA[i])){
namesIncompleteFA[i] = i+1
incompleteKeys = c(incompleteKeys, names$CitationKey[i])
incompleteRows = c(incompleteRows, i+1)
}
namesIncompleteFA = namesIncompleteFA[!is.na(namesIncompleteFA)]
if (nchar(names$LA[i])<2 || grepl("\\.", names$LA[i])){
namesIncompleteLA[i] = i+1
incompleteKeys = c(incompleteKeys, names$CitationKey[i])
incompleteRows = c(incompleteRows, i+1)
}
namesIncompleteLA = namesIncompleteLA[!is.na(namesIncompleteLA)]
}
write.table(incompleteKeys[2:length(incompleteKeys)], "incompleteKeys.csv", sep=",", col.names=FALSE)
write.table(incompleteRows[2:length(incompleteRows)], "incompleteRows.csv", sep=",", col.names=FALSE)
if (length(names$CitationKey[which(names$SelfCite=="Y")]>0)){
print(paste("STOP: Please remove self-citations by searching for the following citation keys in your .bib file: "))
print(paste(names$CitationKey[which(names$SelfCite=="Y")]))
}
if (length(namesIncompleteFA)>0 || length(namesIncompleteLA)>0){
print(paste("STOP: Please revise incomplete full first names or empty cells by searching for the following citation keys in your .bib file: "))
print(paste(incompleteKeys))
print(paste("Do not continue without revising the incomplete names in the citations of your .bib file as indicated above. For more info, see rows", paste(unique(c(namesIncompleteFA, namesIncompleteLA))), "of cleanedBib.csv"))
}
4. Describe the proportions of genders in your reference list and compare it to published base rates in neuroscience.¶
NOTE: your free GenderAPI account has 500 queries per month. This box contains the code that will use your limited API credits/queries if it runs without error. Re-running all code repeatedly will repeatedly use these credits.
Run the code blocks below. (click to select the block and then press Ctrl+Enter; or click the block and press the Run button in the top menubar)
from ethnicolr import pred_fl_reg_name
f = open("genderAPIkey.txt", "r")
genderAPI_key = f.readline().replace('\n', '')
import argparse
parser = argparse.ArgumentParser()
parser.add_argument('-bibfile',action='store',dest='bibfile',default=' '.join(bib_files))
parser.add_argument('-homedir',action='store',dest='homedir',default='/home/jovyan/')
parser.add_argument('-authors',action='store',dest='authors', default=(yourFirstAuthor+' '+yourLastAuthor).replace(',',''))
parser.add_argument('-method',action='store',dest='method',default='florida')
parser.add_argument('-font',action='store',dest='font',default='Palatino') # hey, we all have our favorite
parser.add_argument('-gender_key',action='store',dest='gender_key',default=genderAPI_key)
r = parser.parse_args()
locals().update(r.__dict__)
bibfile = parse_file(bibfile)
def gender_base():
"""
for unknown gender, fill with base rates
you will never / can't run this (that file is too big to share)
"""
main_df = pd.read_csv('/%s/data/NewArticleData2019.csv'%(homedir),header=0)
gender_base = {}
for year in np.unique(main_df.PY.values):
ydf = main_df[main_df.PY==year].AG
fa = np.array([x[0] for x in ydf.values])
la = np.array([x[1] for x in ydf.values])
fa_m = len(fa[fa=='M'])/ len(fa[fa!='U'])
fa_w = len(fa[fa=='W'])/ len(fa[fa!='U'])
la_m = len(la[fa=='M'])/ len(la[la!='U'])
la_w = len(la[fa=='W'])/ len(la[la!='U'])
gender_base[year] = [fa_m,fa_w,la_m,la_w]
gender_base[2020] = [fa_m,fa_w,la_m,la_w]
with open(homedir + '/data/gender_base' + '.pkl', 'wb') as f:
pickle.dump(gender_base, f, pickle.HIGHEST_PROTOCOL)
with open(homedir + 'data/gender_base' + '.pkl', 'rb') as f:
gender_base = pickle.load(f)
authors = authors.split(' ')
print ('first author is %s %s '%(authors[1],authors[0]))
print ('last author is %s %s '%(authors[3],authors[2]))
print ("we don't count these, but check the predictions file to ensure your names did not slip through!")
citation_matrix = np.zeros((8,8))
matrix_idxs = {'white_m':0,'api_m':1,'hispanic_m':2,'black_m':3,'white_f':4,'api_f':5,'hispanic_f':6,'black_f':7}
asian = [0,1,2]
black = [3,4]
white = [5,6,7,8,9,11,12]
hispanic = [10]
print ('looping through your references, predicting gender and race')
columns=['CitationKey','Author','Gender','W','A', 'GendCat']
paper_df = pd.DataFrame(columns=columns)
gender = []
race = []
idx = 0
for paper in tqdm.tqdm(bibfile.entries,total=len(bibfile.entries)):
if 'author' not in bibfile.entries[paper].persons.keys():
continue #some editorials have no authors
if 'year' not in bibfile.entries[paper].fields.keys():
year = 2020
else: year = int(bibfile.entries[paper].fields['year'])
if year not in gender_base.keys():
gb = gender_base[1995]
else:
gb = gender_base[year]
fa = bibfile.entries[paper].persons['author'][0]
try:fa_fname = fa.first_names[0]
except:fa_fname = fa.last_names[0] #for people like Plato
fa_lname = fa.last_names[0]
la = bibfile.entries[paper].persons['author'][-1]
try:la_fname = la.first_names[0]
except:la_fname = la.last_names[0] #for people like Plato
la_lname = la.last_names[0]
if fa_fname.lower().strip() == authors[1].lower().strip():
if fa_lname.lower().strip() == authors[0].lower().strip() :
continue
if fa_fname.lower().strip() == authors[3].lower().strip() :
if fa_lname.lower().strip() == authors[2].lower().strip() :
continue
if la_fname.lower().strip() == authors[1].lower().strip() :
if la_lname.lower().strip() == authors[0].lower().strip() :
continue
if la_fname.lower().strip() == authors[3].lower().strip() :
if la_lname.lower().strip() == authors[2].lower().strip() :
continue
fa_fname = fa_fname.encode("ascii", errors="ignore").decode()
fa_lname = fa_lname.encode("ascii", errors="ignore").decode()
la_fname = la_fname.encode("ascii", errors="ignore").decode()
la_lname = la_lname.encode("ascii", errors="ignore").decode()
names = [{'lname': fa_lname,'fname':fa_fname}]
fa_df = pd.DataFrame(names,columns=['fname','lname'])
asian,hispanic,black,white = pred_fl_reg_name(fa_df,'lname','fname').values[0][-4:]
fa_race = [white,asian,hispanic,black]
names = [{'lname': la_lname,'fname':la_fname}]
la_df = pd.DataFrame(names,columns=['fname','lname'])
asian,hispanic,black,white = pred_fl_reg_name(la_df,'lname','fname').values[0][-4:]
la_race = [white,asian,hispanic,black]
url = "https://gender-api.com/get?key=" + gender_key + "&name=%s" %(fa_fname)
response = urlopen(url)
decoded = response.read().decode('utf-8')
fa_gender = json.loads(decoded)
if fa_gender['gender'] == 'female':
fa_g = [0,fa_gender['accuracy']/100.]
if fa_gender['gender'] == 'male':
fa_g = [fa_gender['accuracy']/100.,0]
if fa_gender['gender'] == 'unknown':
fa_g = gb[:2]
url = "https://gender-api.com/get?key=" + gender_key + "&name=%s" %(la_fname)
response = urlopen(url)
decoded = response.read().decode('utf-8')
la_gender = json.loads(decoded)
if la_gender['gender'] == 'female':
la_g = [0,la_gender['accuracy']/100.]
if la_gender['gender'] == 'male':
la_g = [la_gender['accuracy']/100.,0]
if la_gender['gender'] == 'unknown':
la_g = gb[2:]
fa_data = np.array([paper,'%s,%s'%(fa_fname,fa_lname),'%s,%s'%(fa_gender['gender'],fa_gender['accuracy']),fa_race[0],np.sum(fa_race[1:]), '']).reshape(1,6)
paper_df = paper_df.append(pd.DataFrame(fa_data,columns=columns),ignore_index =True)
la_data = np.array([paper,'%s,%s'%(la_fname,la_lname),'%s,%s'%(la_gender['gender'],la_gender['accuracy']),la_race[0],np.sum(la_race[1:]), '%s%s' % (fa_gender['gender'], la_gender['gender'])]).reshape(1,6)
paper_df = paper_df.append(pd.DataFrame(la_data,columns=columns),ignore_index =True)
mm = fa_g[0]*la_g[0]
wm = fa_g[1]*la_g[0]
mw = fa_g[0]*la_g[1]
ww = fa_g[1]*la_g[1]
mm,wm,mw,ww = [mm,wm,mw,ww]/np.sum([mm,wm,mw,ww])
gender.append([mm,wm,mw,ww])
ww = fa_race[0] * la_race[0]
aw = np.sum(fa_race[1:]) * la_race[0]
wa = fa_race[0] * np.sum(la_race[1:])
aa = np.sum(fa_race[1:]) * np.sum(la_race[1:])
race.append([ww,aw,wa,aa])
paper_matrix = np.zeros((2,8))
paper_matrix[0] = np.outer(fa_g,fa_race).flatten()
paper_matrix[1] = np.outer(la_g,la_race).flatten()
paper_matrix = np.outer(paper_matrix[0],paper_matrix[1])
citation_matrix = citation_matrix + paper_matrix
idx = idx + 1
mm,wm,mw,ww = np.mean(gender,axis=0)*100
WW,aw,wa,aa = np.mean(race,axis=0)*100
statement = "Recent work in several fields of science has identified a bias in citation practices such that papers from women and other minority scholars\
are under-cited relative to the number of such papers in the field (1-5). Here we sought to proactively consider choosing references that reflect the \
diversity of the field in thought, form of contribution, gender, race, ethnicity, and other factors. First, we obtained the predicted gender of the first \
and last author of each reference by using databases that store the probability of a first name being carried by a woman (5, 6). By this measure \
(and excluding self-citations to the first and last authors of our current paper), our references contain ww% woman(first)/woman(last), \
MW% man/woman, WM% woman/man, and MM% man/man. This method is limited in that a) names, pronouns, and social media profiles used to construct the \
databases may not, in every case, be indicative of gender identity and b) it cannot account for intersex, non-binary, or transgender people. \
Second, we obtained predicted racial/ethnic category of the first and last author of each reference by databases that store the probability of a \
first and last name being carried by an author of color (7,8). By this measure (and excluding self-citations), our references contain AA% author of \
color (first)/author of color(last), WA% white author/author of color, AW% author of color/white author, and WW% white author/white author. This method \
is limited in that a) names and Florida Voter Data to make the predictions may not be indicative of racial/ethnic identity, and b) \
it cannot account for Indigenous and mixed-race authors, or those who may face differential biases due to the ambiguous racialization or ethnicization of their names. \
We look forward to future work that could help us to better understand how to support equitable practices in science."
statement = statement.replace('MM',str(np.around(mm,2)))
statement = statement.replace('WM',str(np.around(wm,2)))
statement = statement.replace('MW',str(np.around(mw,2)))
statement = statement.replace('ww',str(np.around(ww,2)))
statement = statement.replace('WW',str(np.around(WW,2)))
statement = statement.replace('AW',str(np.around(aw,2)))
statement = statement.replace('WA',str(np.around(wa,2)))
statement = statement.replace('AA',str(np.around(aa,2)))
5. Print the Diversity Statement and visualize your results¶
The example template can be copied and pasted into your manuscript. We have included it in our methods or references section. If you are using LaTeX, the bibliography file can be found here.
Additional info about the neuroscience benchmark¶
For the top 5 neuroscience journals (Nature Neuroscience, Neuron, Brain, Journal of Neuroscience, and Neuroimage), the expected gender proportions in reference lists as reported by Dworkin et al. are 58.4% for man/man, 9.4% for man/woman, 25.5% for woman/man, and 6.7% for woman/woman. Expected proportions were calculated by randomly sampling papers from 28,505 articles in the 5 journals, estimating gender breakdowns using probabilistic name classification tools, and regressing for relevant article variables like publication date, journal, number of authors, review article or not, and first-/last-author seniority. See Dworkin et al. for more details.
Using a similar random draw model regressing for relevant variables, the expected race proportions in reference lists as reported by Bertolero et al. were 51.8% for white/white, 12.8% for white/author-of-color, 23.5% for author-of-color/white, and 11.9% for author-of-color/author-of-color.
This box does NOT contain code that will use your limited API credits/queries.
Run the code block below. (click to select the block and then press Ctrl+Enter; or click the block and press the Run button in the top menubar)
print (statement)
cmap = sns.diverging_palette(220, 10, as_cmap=True)
names = ['white_m','api_m','hispanic_m','black_m','white_w','api_w','hispanic_w','black_w']
plt.close()
sns.set(style='white')
fig, axes = plt.subplots(ncols=2,nrows=1,figsize=(7.5,4))
axes = axes.flatten()
plt.sca(axes[0])
heat = sns.heatmap(np.around((citation_matrix/citation_matrix.sum())*100,2),annot=True,ax=axes[0],annot_kws={"size": 8},cmap=cmap,vmax=1,vmin=0)
axes[0].set_ylabel('first author',labelpad=0)
heat.set_yticklabels(names,rotation=0)
axes[0].set_xlabel('last author',labelpad=1)
heat.set_xticklabels(names,rotation=90)
heat.set_title('percentage of citations')
citation_matrix_sum = citation_matrix / np.sum(citation_matrix)
expected = np.load('/%s/data/expected_matrix_florida.npy'%(homedir))
expected = expected/np.sum(expected)
percent_overunder = np.ceil( ((citation_matrix_sum - expected) / expected)*100)
plt.sca(axes[1])
heat = sns.heatmap(np.around(percent_overunder,2),annot=True,ax=axes[1],fmt='g',annot_kws={"size": 8},vmax=50,vmin=-50,cmap=cmap)
axes[1].set_ylabel('',labelpad=0)
heat.set_yticklabels('')
axes[1].set_xlabel('last author',labelpad=1)
heat.set_xticklabels(names,rotation=90)
heat.set_title('percentage over/under-citations')
plt.tight_layout()
plt.savefig('/home/jovyan/race_gender_citations.pdf')
paper_df.to_csv('/home/jovyan/predictions.csv')
# Plot a histogram #
names <- read.csv('/home/jovyan/predictions.csv', header=T)
total_citations <- nrow(na.omit(names))
names$GendCat <- gsub("female", "W", names$GendCat, fixed=T)
names$GendCat <- gsub("male", "M", names$GendCat, fixed=T)
names$GendCat <- gsub("unknown", "U", names$GendCat, fixed=T)
gend_cats <- unique(names$GendCat) # get a vector of all the gender categories in your paper
# Create an empty data frame that will be used to plot the histogram. This will have the gender category (e.g., WW, MM) in the first column and the percentage (e.g., number of WW citations divided by total number of citations * 100) in the second column #
dat_for_plot <- data.frame(gender_category = NA,
number = NA,
percentage = NA)
### Loop through each gender category from your paper, calculate the citation percentage of each gender category, and save the gender category and its citation percentage in dat_for_plot data frame ###
if (length(names$GendCat) != 1) {
for (i in 1:length(gend_cats)){
# Create an empty temporary data frame that will be binded to the dat_for_plot data frame
temp_df <- data.frame(gender_category = NA,
number = NA,
percentage = NA)
# Get the gender category, the number of citations with that category, and calculate the percentage of citations with that category
gend_cat <- gend_cats[i]
number_gend_cat <- length(names$GendCat[names$GendCat == gend_cat])
perc_gend_cat <- (number_gend_cat / total_citations) * 100
# Bind this information to the original data frame
temp_df$gender_category <- gend_cat
temp_df$number <- number_gend_cat
temp_df$percentage <- perc_gend_cat
dat_for_plot <- rbind(dat_for_plot, temp_df)
}
}
# Create a data frame with only the WW, MW, WM, MM categories and their base rates - to plot percent citations relative to benchmarks
dat_for_baserate_plot <- subset(dat_for_plot, gender_category == 'WW' | gender_category == 'MW' | gender_category == 'WM' | gender_category == 'MM')
dat_for_baserate_plot$baserate <- c(6.7, 9.4, 25.5, 58.4)
dat_for_baserate_plot$citation_rel_to_baserate <- dat_for_baserate_plot$percentage - dat_for_baserate_plot$baserate
# Plot the Histogram of Number of Papers per category against predicted gender category #
library(ggplot2)
dat_for_plot = dat_for_plot[-1:-2,]
dat_for_plot$gender_category <- factor(dat_for_plot$gender_category, levels = dat_for_plot$gender_category)
ggplot(dat_for_plot[-c(1),], aes(x = gender_category, y = number, fill = gender_category)) +
geom_bar(stat = 'identity', width = 0.75, na.rm = TRUE, show.legend = TRUE) +
scale_x_discrete(limits = c('WW', 'MW', 'WM', 'MM', 'UW', 'UM', 'WU', 'MU', 'UU')) +
geom_text(aes(label = number), vjust = -0.3, color = 'black', size = 2.5) +
theme(legend.position = 'right') + theme_minimal() +
xlab('Predicted gender category') + ylab('Number of papers') + ggtitle("") + theme_classic(base_size=15)
# Plot the Histogram of % citations relative to benchmarks against predicted gender category
ggplot(dat_for_baserate_plot, aes(x = gender_category, y = citation_rel_to_baserate, fill = gender_category)) +
geom_bar(stat = 'identity', width = 0.75, na.rm = TRUE, show.legend = TRUE) +
scale_x_discrete(limits = c('WW', 'MW', 'WM', 'MM')) +
geom_text(aes(label = round(citation_rel_to_baserate, digits = 2)), vjust = -0.3, color = 'black', size = 2.5) +
theme(legend.position = 'right') + theme_minimal() +
xlab('Predicted gender category') + ylab('% of citations relative to benchmarks') + ggtitle("") + theme_classic(base_size=15)
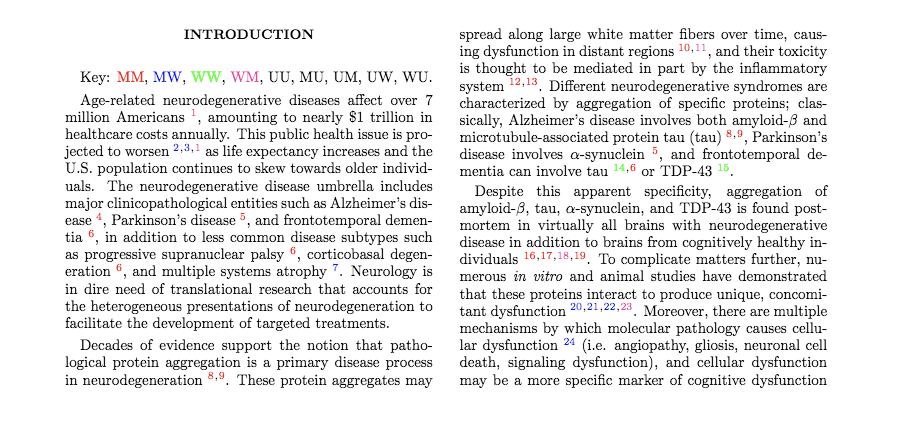
(OPTIONAL) Color-code your .tex file using the estimated gender classifications¶
Running this code-block will optionally output your uploaded .tex file with color-coding for gender pair classifications. You can find the example below's pre-print here.

cite_gender = pd.read_csv(homedir+'Authors.csv') # output of getReferenceGends.ipynb
cite_gender.index = cite_gender.CitationKey
cite_gender['Color'] = '' # what color to make each gender category
colors = {'MM':'red','MW':'blue','WW':'green','WM':'magenta','UU':'black',
'MU':'black','UM':'black','UW':'black','WU':'black'}
for idx in cite_gender.index: # loop through each citation key and set color
cite_gender.loc[idx,'Color'] = colors[cite_gender.loc[idx,'GendCat']]
cite_gender.loc[cite_gender.index[cite_gender.SelfCite=='Y'],'Color'] = 'black' # make self citations black
fin = open(homedir+tex_file)
texdoc=fin.readlines()
with open(homedir+tex_file[:-4]+'_gendercolor.tex','w') as fout:
for i in range(len(texdoc)):
s = texdoc[i]
cite_instances = re.findall('\\\\cite\{.*?\}',s)
cite_keys = re.findall('\\\\cite\{(.*?)\}',s)
cite_keys = [x.split(',') for x in cite_keys]
cite_keys_sub = [['\\textcolor{' + cite_gender.loc[x.strip(),'Color'] + '}{\\cite{'+x.strip()+'}}' for x in cite_instance] for cite_instance in cite_keys]
cite_keys_sub = ['\\textsuperscript{,}'.join(x) for x in cite_keys_sub]
for idx,cite_instance in enumerate(cite_instances):
s = s.replace(cite_instances[idx],cite_keys_sub[idx])
fout.write(s)
# place color key after abstract
if '\\section*{Introduction}\n' in s:
l = ['\\textcolor{' + colors[k] + '}{'+k+'}' for k in colors.keys()]
fout.write('\tKey: '+ ', '.join(l)+'.\n')