Machine Learning and Statistics for Physicists¶
Material for a UC Irvine course offered by the Department of Physics and Astronomy.
Content is maintained on github and distributed under a BSD3 license.
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import numpy as np
import pandas as pd
We will use the sklearn decomposition module below:
from sklearn import decomposition
Load Data¶
from mls import locate_data
line_data = pd.read_csv(locate_data('line_data.csv'))
pong_data = pd.read_hdf(locate_data('pong_data.hf5'))
cluster_3d = pd.read_hdf(locate_data('cluster_3d_data.hf5'))
cosmo_targets = pd.read_hdf(locate_data('cosmo_targets.hf5'))
spectra_data = pd.read_hdf(locate_data('spectra_data.hf5'))
Data Dimensionality¶
We call the number of features (columns) in a dataset its "dimensionality". In order to learn how different features are related, we need enough samples to get a complete picture.
For example, imagine splitting each feature at its median value then, at a minimum, we would like to have at least one sample in each of the resulting $2^D$ bins (D = dimensionality = # of features = # of columns). This is a very low bar and only requires 8 samples with D=3, but requires $2^{30} > 1$ billion samples with D=30.
To get a feel for how well sampled your dataset is, estimate how many bins you could split each feature (axis) into and end up with 1 sample per bin (assuming that features are uncorrelated). A value < 2 fails our minimal test above and anything < 5 is a potential red flag.
stats = []
for name in 'line_data', 'cluster_3d', 'cosmo_targets', 'pong_data', 'spectra_data':
N, D = eval(name).shape
stats.append([name, N, D, N ** (1 / D)])
pd.DataFrame(stats, columns=('Dataset', 'N', 'D', 'N**(1/D)')).round(3)
| Dataset | N | D | N**(1/D) | |
|---|---|---|---|---|
| 0 | line_data | 2000 | 3 | 12.599 |
| 1 | cluster_3d | 500 | 3 | 7.937 |
| 2 | cosmo_targets | 50000 | 6 | 6.070 |
| 3 | pong_data | 1000 | 20 | 1.413 |
| 4 | spectra_data | 200 | 500 | 1.011 |
However, not all features carry equal information and the effective dimensionality of a dataset might be lower than the number of columns. As an extreme example, consider the following 2D data which is effectively 1D since one column has a constant value (zero):
gen = np.random.RandomState(seed=123)
data = pd.DataFrame()
data['x'] = gen.uniform(-1, +1, 50)
data['y'] = np.zeros_like(data['x'])
sns.jointplot(x='x', y='y', data=data);
DISCUSS: Is this data is still 1D if (refer to the plots below):
- we add some small scatter in the 2nd dimension?
- we perform a coordinate rotation so that y ~ m x?
- y ~ f(x) where f(x) is nonlinear?
The scatter adds new information in a second dimension, but we can approximately ignore it under two assumptions:
- The relative scaling of the x and y columns is meaningful (which is almost certainly not true if these columns have different dimensions - recall our earlier comments about normalizing data).
- The origin of the scatter is due to measurement error or some other un-informative process.
The rotation does not change the effective dimensionality of the data.
A non-linear relationship between x and y also does not change the underlying dimensionality since we could, in principle, perform a non-linear change of coordinates to undo it. However, we can expect that non-linear relationships will be harder to deal with than linear ones.
# Add some scatter in the 2nd dimension.
data['y'] = gen.normal(scale=0.04, size=len(data))
sns.jointplot(x='x', y='y', data=data);
plt.ylim(-1, +1);
# Rotate by 30 deg counter-clockwise.
theta = np.deg2rad(30.)
rotated = data.copy()
rotated['x'] = np.cos(theta) * data['x'] - np.sin(theta) * data['y']
rotated['y'] = np.cos(theta) * data['x'] + np.sin(theta) * data['y']
sns.jointplot(x='x', y='y', data=rotated);
# Use the nonlinear y ~ x ** 2 + x instead of y ~ x.
nonlinear = rotated.copy()
nonlinear['y'] = rotated['y'] + rotated['x'] ** 2
sns.jointplot(x='x', y='y', data=nonlinear);
We will use spectra_data below. Note from the table above that it appears to be severely undersampled with N=200, D=500.
EXERCISE: Plot some rows (samples) of spectra_data using plt.plot(spectra_data.iloc[i], '.') to get a feel for this dataset. What do you think the effective dimensionality of this data is? (Hint: how many independent parameters you would need to generate this data?)
for i in range(10):
plt.plot(spectra_data.iloc[i], '.')
Each sample is a graph of a smooth function with some noise added. The smooth function has three distinct components:
- two peaks, with fixed locations and shapes, and normalizations that vary independently.
- a smooth background with no free parameters.
Since the data could be reproduced with just normalization parameters (except for the noise), it has an effective dimensionality of $d=2$.
Note that the relative normalization of each feature is significant here, so we would not want to normalize this data and lose this information. We refer to each sample as a "spectrum" since it looks similar to spectra obtained in different areas of physics (astronomy, nuclear physics, particle physics, ...)
# Add your solution here...
Linear Decompositions¶
The goal of a linear decomposition is to automatically identify linear combinations of the original features that account for most of the variance in the data. Note that we are using variance (spread) as a proxy for "useful information", so it is essential that the relative normalization of our features is meaningful.
If we represent our data with the $N\times D$ matrix $X$, then a linear decomposition can be represented as the following matrix multiplication:
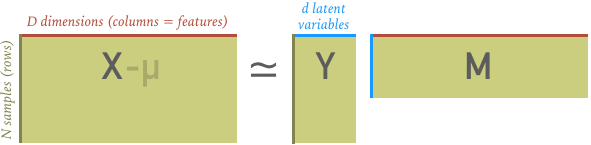
The $N\times d$ matrix $Y$ is a reduced representation of the original data $X$, with $d < D$ new features that are linear combinations of the original $D$ feature. We call the new features "latent variables", since they were already present in $X$ but only implicitly.
The $d\times D$ matrix $M$ specifies the relationship between the old and new features: each column is a unit vector for a new feature in terms of the old features. Note that $M$ is not square when $d < D$ and unit vectors are generally not mutually orthogonal (except for the PCA method).
A linear decomposition is not exact (hence the $\simeq$ above) and there is no "best" prescription for determining $Y$ and $M$. Below we review the most popular prescriptions implemented in the sklearn.decomposition module (links are to wikipedia and sklearn documentation):
| Method | sklearn |
|---|---|
| Principal Component Analysis | PCA |
| Factor Analysis | FactorAnalysis |
| Non-negative Matrix Factorization | NMF |
| Independent Component Analysis | FastICA |
All methods require that you specify the number of latent variables $d$ (but you can easily experiment with different values) and are called using (method = PCA, FactorAnalysis, NMF, FastICA):
fit = decomposition.method(n_components=d).fit(X)
The resulting decomposition into $Y$ and $M$ is given by:
M = fit.components_
Y = fit.transform(X)
except for FastICA, where M = fit.mixing_.T.
When $d < D$, we refer to the decomposition as a "dimensionality reduction". A useful visualization of how effectively the latent variables capture the interesting information in the original data is to calculate: $$ X' = Y M \; , $$ which approximately reconstructs the original data, and compare rows (samples) of $X'$ with the original $X$. They will not agree exactly, but if the differences seem uninteresting (e.g., look like noise), then the dimensionality reduction was successful and you can use $Y$ instead of $X$ for subsequent analysis.
We will use the function below to demonstrate each of these in turn (but you can ignore its details unless you are interested):
def demo(method='PCA', d=2, data=spectra_data):
X = data.values
N, D = X.shape
if method == 'NMF':
# All data must be positive.
assert np.all(X > 0)
# Analysis includes the mean.
mu = np.zeros(D)
else:
mu = np.mean(X, axis=0)
kwargs = dict(n_components=d)
if method == 'NMF':
kwargs['max_iter'] = 500
fit = eval('decomposition.' + method)(**kwargs).fit(X)
# Check that decomposition has the expected shape.
if method == 'FastICA':
M = fit.mixing_.T
else:
M = fit.components_
assert M.shape == (d, D)
Y = fit.transform(X)
assert Y.shape == (N, d)
# Reconstruct X - mu from the fitted Y, M.
Xr = np.dot(Y, M) + mu
assert Xr.shape == X.shape
# Plot pairs of latent vars.
columns = ['y{}'.format(i) for i in range(d)]
sns.pairplot(pd.DataFrame(Y, columns=columns))
plt.show()
# Compare a few samples from X and Xr.
fig = plt.figure(figsize=(8.5, 4))
for i,c in zip((0, 6, 7), sns.color_palette()):
plt.plot(X[i], '.', c=c, ms=3, alpha=0.5)
plt.plot(Xr[i], '-', c=c, lw=2)
plt.xlim(-0.5, D+0.5)
plt.xlabel('Feature #')
plt.ylabel('Feature Value')
label = '{}(d={}): $\sigma = {:.2f}$'.format(method, d, np.std(Xr - X))
plt.text(0.95, 0.9, label, horizontalalignment='right',
fontsize='x-large', transform=plt.gca().transAxes)
Principal Component Analysis¶
PCA is the most commonly used method for dimensionality reduction. The decomposition is uniquely specified by the following prescription (more details here):
- Find the eigenvectors and eigenvalues of
which is an empirical estimate of the covariance matrix for the data.
- Construct $M$ out of the eigenvectors, ordered by decreasing eigenvalue (which are all positive) and solve the resulting linear equations for $Y$. At this point the decomposition is exact with $d = D$.
- Shrink $Y$ and $M$ from $D$ to $d$ rows ($M$) or columns ($Y$), which makes the decomposition approximate while discarding the least amount of variance in the original data (which we use as a proxy for "useful information").
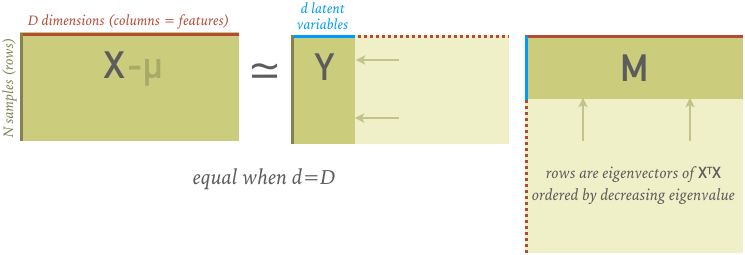
The full $M$ matrix (before shrinking $D\rightarrow d$) is orthogonal $M^T = M^{-1}$ and satisfies $X^T X = M^T \Lambda M$, where $\Lambda$ is a diagonal matrix of the decreasing eigenvalues. Note that this description glosses over some details that you will explore in your homework.
The resulting latent variables are statistically uncorrelated (which is a weaker statement than statistically independent -- see below), i.e., the correlation coefficients between different columns of $Y$ are approximately zero: $$ \rho(j,k) = \frac{Y_j\cdot Y_k}{|Y_j|\,|Y_k|} \simeq 0 \; . $$
The PCA demonstration below shows a pairplot of the latent variables from a $d=2$ decomposition, followed by a reconstruction of some samples (red curves) compared with the originals (red points).
Note that the reconstructed samples are in some sense better than the originals since the original noise was associated with a small eigenvalue that was trimmed!
demo('PCA', d=2)
DISCUSS: How many clusters do you expect to see in the scatter plot of y0 versus y1 above based on what you know about this dataset? Can you identify these clusters in plot above?
We expect to see 4 clusters, corresponding to spectra with:
- No peaks.
- Only the lower peak.
- Only the upper peak.
- Both peaks.
We already saw that this data can be generated from two flux values, giving the normalization of each peak. Lets assume that y0 and y1 are related to these fluxes to identify the clusters:
- Points near (-2000, -2000), with very little spread.
- Points along the horizontal line with y0 ~ -2000.
- Points along the diagonal line.
- Points scattered between the two lines.
Factor Analysis¶
Factor analysis (FA) often produces similar results to PCA, but is conceptually different.
Both PCA and FA implicitly assume that the data is approximately sampled from a multidimensional Gaussian. PCA then finds the principal axes of the the resulting multidimensional ellipsoid, while FA is based on a model for how the original data is generated from the latent variables. Specifically, FA seeks latent variables that are uncorrelated unit Gaussians and allows for different noise levels in each feature, while assuming that this noise is uncorrelated with the latent variables. PCA does not distinguish between "signal" and "noise" and implicitly assumes that the large eigenvalues are more signal-like and small ones more noise-like.
When the FA assumptions about the data (of Gaussian latent variables with uncorrelated noise added) are correct, it is certaintly the better choice, in principle. In practice, FA decomposition is more expensive and requires an iterative Expectation-Maximization (EM) algorithm. You should normally try both, but prefer the simpler PCA when the results are indistinguishable.
demo('FactorAnalysis', d=2)
Non-negative Matrix Factorization¶
Most linear factorizations start by centering each feature about its mean over the samples: $$ X_{ij} \rightarrow X_{ij} - \mu_i \quad , \quad \mu_i \equiv \frac{1}{N} \sum_i\, X_{ij} \; . $$ As a result, latent variables are equally likely to be positive or negative.
Non-negative matrix factorization (NMF) assumes that the data are a (possibly noisy) linear superposition of different components, which is often a good description of data resulting from a physical process. For example, the spectrum of a galaxy is a superposition of the spectra of its constituent stars, and the spectrum of a radioactive sample is a superposition of the decays of its constituent unstable isotopes.
These processes can only add data, so the elements of $Y$ and $M$ should all be $\ge 0$ if the latent variables describe additive mixtures of different processes. The NMF factorization guarantees that both $Y$ and $M$ are positive, and requires that the input $X$ is also positive. However, there is no guarantee that the non-negative latent variables found by NMF are due to physical mixtures.
Since NMF does not internally subtract out the means $\mu_i$, it generally needs an additional component to model the mean. For spectra_data then, we should use d=3 for NMF to compare with PCA using d=2:
demo('NMF', d=3)
To see the importance of the extra latent variable, try with d=2 and note how poorly samples are reconstructed:
demo('NMF', d=2)
Idependent Component Analysis¶
The final linear decomposition we will consider is ICA, where the goal is to find latent variables $Y$ that are statistically independent, which is a stronger statement that the statistically uncorrelated guarantee of PCA. We will formalize the definition of independence soon but the basic idea is that the joint probability of a sample occuring with latent variables $y_1, y_2, y_3, \ldots$ can be factorized into independent probabilities for each component: $$ P(y_1, y_2, y_3, \ldots) = P(y_1) P(y_2) P(y_3) \ldots $$
ICA has some inherent ambiguities: both the ordering and scaling of latent variables is arbitrary, unlike with PCA. However, in practice, samples reconstructed with ICA often look similar to PCA and FA reconstructions.
ICA is also used for blind signal separation, where usually $d = N$.
demo('FastICA', d=2)
Comparison of Linear Methods¶
To compare the four methods above, plot their normalized "unit vectors" (rows of the $M$ matrix). Note that only the NMF curves are always positive, as expected. However, while all methods give excellent reconstructions of the original data, they also all mix the two peaks together. In other words, none of the methods has discovered the natural latent variables of the underlying physical process: the individual peak normalizations.
def compare_linear(data=spectra_data):
X = data.values
N, D = X.shape
fig = plt.figure(figsize=(8.5, 4))
for j, method in enumerate(('PCA', 'FactorAnalysis', 'NMF', 'FastICA')):
if method == 'NMF':
d = 3
mu = np.zeros(D)
else:
d = 2
mu = np.mean(X, axis=0)
kwargs = dict(n_components=d)
if method == 'NMF':
kwargs['max_iter'] = 500
fit = eval('decomposition.' + method)(**kwargs).fit(X - mu)
M = fit.mixing_.T if method == 'FastICA' else fit.components_
for i in range(d):
unitvec = M[i] / M[i].sum()
plt.plot(unitvec, label=method, c=sns.color_palette()[j], ls=('-', '--', ':')[i])
plt.legend()
plt.xlim(-0.5, D + 0.5)
compare_linear()
Weighted PCA¶
The linear algorithms presented above work fine with noisy data, but have no way to take advantage of data that includes its own estimate of the noise level. In the most general case, each element of the data matrix $X$ has a corresponding RMS error estimate $\delta X$, with values $\rightarrow\infty$ used to indicate missing data. In practice, it is convenient to replace $\delta X$ with a matrix of weights $W$ where zero values indicate missing data. For data with Gaussian errors, $X_{ij} \pm \delta X_{ij}$, the appropriate weight is usually the inverse variance $I \equiv \delta X_{ij}^{-2}$.
Delchambre 2015 reviews several approaches for calculating principal components of weighted data, and proposes a new scheme where, instead of subtracting the usual mean, we subtract a weighted mean, $$ \mu_j = \frac{\sum_{i=1}^N W_{ij} X_{ij}}{\sum_{i=1}^N W_{ij}} \; , $$ with $W = \sqrt{I}$, and, instead of decomposing $C = X^T X$, we decompose $$ C = \frac{(W X)^T (W X)}{W^T W} \; . $$
The class below implements this scheme following the same API as the sklearn PCA class:
# Use scipy.linalg.eigh instead of np.linalg.eigh since it allows us to only
# find the eigenvectors with the largest eigenvalues.
import scipy.linalg
class WeightedPCA(object):
"""Implements the weighted PCA scheme of Delchambre 2015.
See https://arxiv.org/abs/1412.4533 and the more complete implementation
in https://github.com/jakevdp/wpca.
"""
def __init__(self, n_components):
self.n_components = n_components
def prepare(self, X, ivar=None):
if ivar is None:
W = np.ones_like(X)
else:
assert np.all(ivar >= 0)
W = np.sqrt(ivar)
# Calculate the weighted mean of the data using eqn (2).
self.mu = np.sum(W * X, axis=0) / np.sum(W, axis=0)
# Subtract the weighted mean from the data.
X = X - self.mu
# Apply weights to the (mean subtracted) data.
X *= W
return X, W
def fit(self, X, ivar=None):
X, W = self.prepare(X, ivar)
# Calculate the weighted covariance.
C = np.dot(X.T, X)
C /= np.dot(W.T, W)
# Find the eigenvectors and eigenvalues of C.
_, D = X.shape
evals, evecs = scipy.linalg.eigh(C, eigvals=(D - self.n_components, D - 1))
# Save the results.
self.components_ = evecs[:, ::-1].T
return self
def transform(self, X, ivar=None):
X, W = self.prepare(X, ivar)
N, _ = X.shape
Y = np.zeros((N, self.n_components))
for i in range(N):
cW = self.components_ * W[i]
cWX = np.dot(cW, X[i])
cWc = np.dot(cW, cW.T)
Y[i] = np.linalg.solve(cWc, cWX)
return Y
def inverse_transform(self, X):
return self.mu + np.dot(X, self.components_)
To study these schemes, we will assign weights to spectra_data by assuming that each value $X_{ij}$ is the result of a Poisson process so has inverse variance $I = X_{ij}^{-1}$.
The function below allows us to optionally add extra noise that varies across the spectra and remove random chunks of data (by setting their weights to zero). As usual, ignore the details of this function unless you are interested.
def weighted_pca(d=2, add_noise=None, missing=None, data=spectra_data, seed=123):
gen = np.random.RandomState(seed=seed)
X = data.values.copy()
N, D = X.shape
# Calculate inverse variances assuming Poisson fluctuations in X.
ivar = X ** -1
# Add some Gaussian noise with a linearly varying RMS, if requested.
if add_noise:
start, stop = add_noise
assert start >= 0 and stop >= 0
extra_rms = np.linspace(start, stop, D)
ivar = (X + extra_rms ** 2) ** -1
X += gen.normal(scale=extra_rms, size=X.shape)
# Remove some fraction of data from each sample, if requested.
if missing:
assert 0 < missing < 0.5
start = gen.uniform(high=(1 - missing) * D, size=N).astype(int)
stop = (start + missing * D).astype(int)
for i in range(N):
X[i, start[i]:stop[i]] = ivar[i, start[i]:stop[i]] = 0.
# Perform the fit.
fit = WeightedPCA(n_components=d).fit(X, ivar)
Y = fit.transform(X, ivar)
Xr = fit.inverse_transform(Y)
# Show the reconstruction of some samples.
fig = plt.figure(figsize=(8.5, 4))
for i,c in zip((0, 6, 7), sns.color_palette()):
plt.plot(X[i], '.', c=c, ms=3, alpha=0.5)
plt.plot(Xr[i], '-', c=c, lw=2)
plt.xlim(-0.5, D+0.5)
plt.xlabel('Feature #')
plt.ylabel('Feature Value')
First check that we recover similar results to the standard PCA with no added noise or missing data. The results are not identical (but presumably better now) because we are accounting for the fact that the relative errors are larger for lower fluxes.
weighted_pca(d=2)
Next, vary the noise level over the spectrum. The larger errors makes the principal components harder to nail down, leading to noisier reconstructions:
weighted_pca(d=2, add_noise=(0, 100))
weighted_pca(d=2, add_noise=(200, 100))
Finally, remove 10% of the data from each sample (but, crucially, a different 10% from each sample). Note how this allows us to make a sensible guess at the missing data! (statisticians call this imputation.
weighted_pca(d=2, missing=0.1, seed=2)
Non-Linear Dimensionality Reduction¶
The methods above find latent variables that are linear functions of the original features.
There are also non-linear methods:
- When they work, the results are spectacular (see also here).
- However, they are generally very sensitive to your choice of hyperparameters.
- I recommend that you always start with a linear method and only prefer a non-linear model if it has clearly better performance and gives consistent and robust results.
For a deeper dive into non-linear methods, see this notebook. One key idea is the "kernel trick", which is also central to the power of neural networks.
Model-Driven Compression¶
Most dimensionality reduction methods assume that variance is a good proxy for "information". However, when you have a good generative model of your data, you can do much better. In particular, there is an optimal compression algorithm for data generated by a model with $n$ parameters that will reduce your whole dataset down to $n$ numbers! See this 2018 paper for details.