Minimizing Kullback-Leibler Divergence¶
In this post, we will see how the KL divergence can be computed between two distribution objects, in cases where an analytical expression for the KL divergence is known. This is the summary of lecture "Probabilistic Deep Learning with Tensorflow 2" from Imperial College London.
- toc: true
- badges: true
- comments: true
- author: Chanseok Kang
- categories: [Python, Coursera, Tensorflow_probability, ICL]
- image: images/kl_pq.gif
Packages¶
In [1]:
import tensorflow as tf
import tensorflow_probability as tfp
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse
from IPython.display import HTML, Image
tfd = tfp.distributions
tfpl = tfp.layers
tfb = tfp.bijectors
plt.rcParams['figure.figsize'] = (10, 6)
plt.rcParams["animation.html"] = "jshtml"
plt.rcParams['animation.embed_limit'] = 2**128
In [2]:
print("Tensorflow Version: ", tf.__version__)
print("Tensorflow Probability Version: ", tfp.__version__)
Tensorflow Version: 2.5.0 Tensorflow Probability Version: 0.13.0
Overview¶
KL divergence¶
In [3]:
# KL(q || p) = E_{z ~ q}[log q(z) - log p(z)]
scale_tril = tfb.FillScaleTriL()([-0.5, 1.25, 1.])
scale_tril
Out[3]:
<tf.Tensor: shape=(2, 2), dtype=float32, numpy=
array([[1.3132716, 0. ],
[1.25 , 0.474087 ]], dtype=float32)>
In [4]:
p = tfd.MultivariateNormalTriL(loc=0., scale_tril=scale_tril)
p
Out[4]:
<tfp.distributions.MultivariateNormalTriL 'MultivariateNormalTriL' batch_shape=[] event_shape=[2] dtype=float32>
In [5]:
q = tfd.MultivariateNormalDiag(loc=[0., 0.])
q
Out[5]:
<tfp.distributions.MultivariateNormalDiag 'MultivariateNormalDiag' batch_shape=[] event_shape=[2] dtype=float32>
In [6]:
tfd.kl_divergence(q, p)
Out[6]:
<tf.Tensor: shape=(), dtype=float32, numpy=3.056092>
Another example¶
In [7]:
q = tfd.MultivariateNormalDiag(
loc=tf.Variable(tf.random.normal([2])),
scale_diag=tfp.util.TransformedVariable(tf.random.uniform([2]), bijector=tfb.Exp())
)
q
Out[7]:
<tfp.distributions.MultivariateNormalDiag 'MultivariateNormalDiag' batch_shape=[] event_shape=[2] dtype=float32>
In [8]:
tfd.kl_divergence(q, p)
Out[8]:
<tf.Tensor: shape=(), dtype=float32, numpy=2.8571239>
In [9]:
@tf.function
def loss_and_grads(q_dist):
with tf.GradientTape() as tape:
loss = tfd.kl_divergence(q_dist, p)
return loss, tape.gradient(loss, q_dist.trainable_variables)
In [10]:
optimizer = tf.keras.optimizers.Adam()
for i in range(20):
loss, grads = loss_and_grads(q)
optimizer.apply_gradients(zip(grads, q.trainable_variables))
print(loss)
tf.Tensor(2.8571239, shape=(), dtype=float32) tf.Tensor(2.8537352, shape=(), dtype=float32) tf.Tensor(2.8503537, shape=(), dtype=float32) tf.Tensor(2.84698, shape=(), dtype=float32) tf.Tensor(2.8436131, shape=(), dtype=float32) tf.Tensor(2.840254, shape=(), dtype=float32) tf.Tensor(2.8369021, shape=(), dtype=float32) tf.Tensor(2.8335583, shape=(), dtype=float32) tf.Tensor(2.8302217, shape=(), dtype=float32) tf.Tensor(2.8268933, shape=(), dtype=float32) tf.Tensor(2.8235722, shape=(), dtype=float32) tf.Tensor(2.8202596, shape=(), dtype=float32) tf.Tensor(2.8169546, shape=(), dtype=float32) tf.Tensor(2.8136582, shape=(), dtype=float32) tf.Tensor(2.81037, shape=(), dtype=float32) tf.Tensor(2.8070896, shape=(), dtype=float32) tf.Tensor(2.8038177, shape=(), dtype=float32) tf.Tensor(2.8005545, shape=(), dtype=float32) tf.Tensor(2.7972991, shape=(), dtype=float32) tf.Tensor(2.7940533, shape=(), dtype=float32)
Tutorial¶
In [11]:
# Define a target distribution, p
tf.random.set_seed(41)
p_mu = [0., 0.]
p_L = tfb.Chain([tfb.TransformDiagonal(tfb.Softplus()),
tfb.FillTriangular()])(tf.random.uniform([3]))
p = tfd.MultivariateNormalTriL(loc=p_mu, scale_tril=p_L)
p
Out[11]:
<tfp.distributions.MultivariateNormalTriL 'MultivariateNormalTriL' batch_shape=[] event_shape=[2] dtype=float32>
In [12]:
# Plot the target distribution's density contours
def plot_density_contours(density, X1, X2, contour_kwargs, ax=None):
'''
Plots the contours of a bivariate TensorFlow density function (i.e. .prob()).
X1 and X2 are numpy arrays of mesh coordinates.
'''
if ax==None:
_, ax = plt.subplots(figsize=(7, 7))
X = np.hstack([X1.flatten()[:, np.newaxis], X2.flatten()[:, np.newaxis]])
density_values = np.reshape(density(X).numpy(), newshape=X1.shape)
ax.contour(X1, X2, density_values, **contour_kwargs)
return(ax)
x1 = np.linspace(-5, 5, 1000)
x2 = np.linspace(-5, 5, 1000)
X1, X2 = np.meshgrid(x1, x2)
f, ax = plt.subplots(1, 1, figsize=(7, 7))
# Density contours are linearly spaced
contour_levels = np.linspace(1e-4, 10**(-0.8), 20) # specific to this seed
ax = plot_density_contours(p.prob, X1, X2,
{'levels':contour_levels,
'cmap':'cividis'}, ax=ax)
ax.set_xlim(-5, 5); ax.set_ylim(-5, 5);
ax.set_title('Density contours of target distribution, $p$')
ax.set_xlabel('$x_1$'); ax.set_ylabel('$x_2$')
plt.show()
In [13]:
# Instantiate an approximating distribution, q, that has diagonal covariance
tf.random.set_seed(41)
q = tfd.MultivariateNormalDiag(loc=tf.Variable(tf.random.normal([2])),
scale_diag=tfp.util.TransformedVariable(tf.random.uniform([2]),
bijector=tfb.Exp()))
q
Out[13]:
<tfp.distributions.MultivariateNormalDiag 'MultivariateNormalDiag' batch_shape=[] event_shape=[2] dtype=float32>
In [14]:
# Define a function for the Kullback-Leibler divergence
@tf.function
def loss_and_grads(dist_a, dist_b):
with tf.GradientTape() as tape:
loss = tfd.kl_divergence(dist_a, dist_b)
return loss, tape.gradient(loss, dist_a.trainable_variables)
In [15]:
from matplotlib import animation
In [16]:
# Run a training loop that computes KL[q || p], updates q's parameters using its gradients
fig, ax1 = plt.subplots(figsize=(7, 7))
num_train_steps = 250
opt = tf.keras.optimizers.Adam(learning_rate=.01)
last_q_loss = 0
def animate(i):
ax1.clear()
global last_q_loss
# Compute the KL divergence and its gradients
q_loss, grads = loss_and_grads(q, p)
# Update the trainable variables using the gradients via the optimizer
opt.apply_gradients(zip(grads, q.trainable_variables))
X = np.hstack([X1.flatten()[:, np.newaxis], X2.flatten()[:, np.newaxis]])
density_values = np.reshape(p.prob(X).numpy(), newshape=X1.shape)
ax1.contour(X1, X2, density_values, levels=contour_levels, cmap='cividis', alpha=0.5)
density_values = np.reshape(q.prob(X).numpy(), newshape=X2.shape)
ax1.contour(X1, X2, density_values, levels=contour_levels, cmap='plasma')
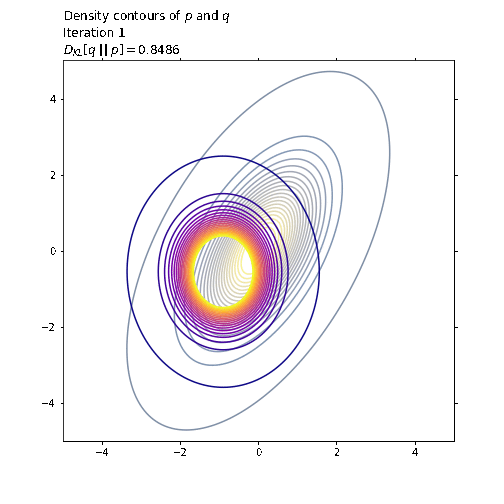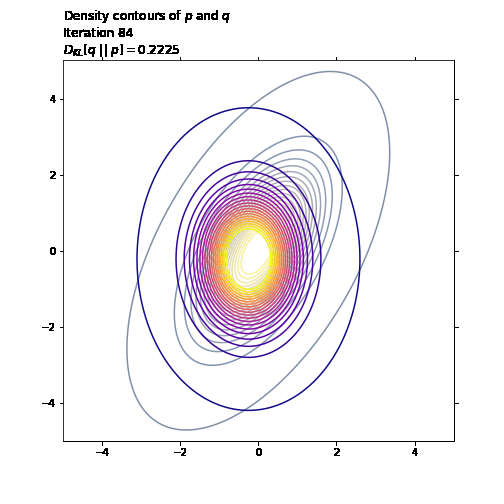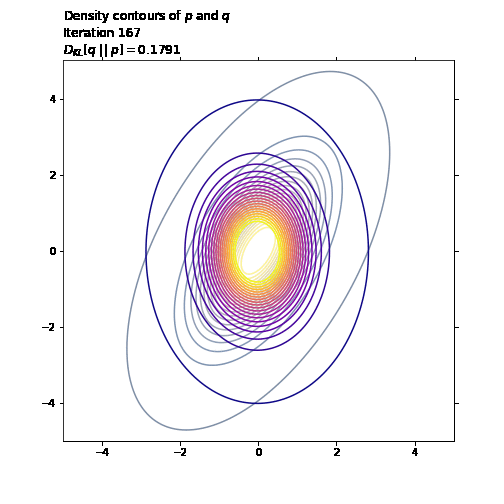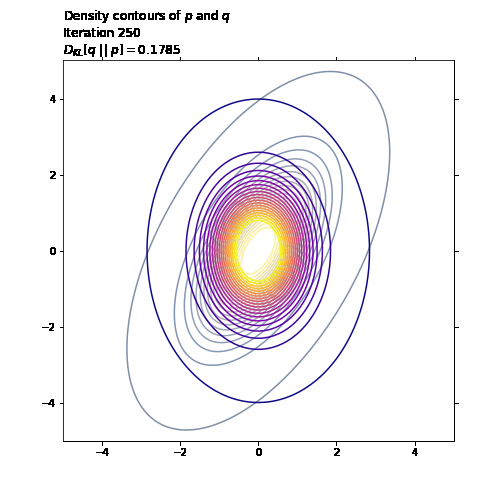
ax1.set_title('Density contours of $p$ and $q$\n' +
'Iteration ' + str(i + 1) + '\n' +
'$D_{KL}[q \ || \ p] = ' +
str(np.round(q_loss.numpy(), 4)) + '$',
loc='left')
last_q_loss = q_loss.numpy()
ani = animation.FuncAnimation(fig, animate, frames=num_train_steps)
plt.close()
In [17]:
ani.save('./image/kl_qp.gif', writer='imagemagick', fps=30)
In [18]:
# Re-fit the distribution, this time fitting q_rev by minimizing KL[p || q_rev]
tf.random.set_seed(41)
q_rev = tfd.MultivariateNormalDiag(loc=tf.Variable(tf.random.normal([2])),
scale_diag=tfp.util.TransformedVariable(tf.random.uniform([2]), bijector=tfb.Exp()))
q_rev
Out[18]:
<tfp.distributions.MultivariateNormalDiag 'MultivariateNormalDiag' batch_shape=[] event_shape=[2] dtype=float32>
In [19]:
# Edit loss_and_grads function
@tf.function
def loss_and_grads(dist_a, dist_b, reverse=False):
with tf.GradientTape() as tape:
if not reverse:
loss = tfd.kl_divergence(dist_a, dist_b)
else:
loss = tfd.kl_divergence(dist_b, dist_a)
return loss, tape.gradient(loss, dist_a.trainable_variables)
In [20]:
# Run a training loop that computes KL[q || p], updates q's parameters using its gradients
fig, ax1 = plt.subplots(figsize=(7, 7))
num_train_steps = 250
opt = tf.keras.optimizers.Adam(learning_rate=.01)
last_q_rev_loss = 0
def animate(i):
ax1.clear()
global last_q_rev_loss
# Compute the KL divergence and its gradients
q_rev_loss, grads = loss_and_grads(q_rev, p, reverse=True)
# Update the trainable variables using the gradients via the optimizer
opt.apply_gradients(zip(grads, q_rev.trainable_variables))
X = np.hstack([X1.flatten()[:, np.newaxis], X2.flatten()[:, np.newaxis]])
density_values = np.reshape(p.prob(X).numpy(), newshape=X1.shape)
ax1.contour(X1, X2, density_values, levels=contour_levels, cmap='cividis', alpha=0.5)
density_values = np.reshape(q_rev.prob(X).numpy(), newshape=X2.shape)
ax1.contour(X1, X2, density_values, levels=contour_levels, cmap='plasma')
ax1.set_title('Density contours of $p$ and $q_{rev}$\n' +
'Iteration ' + str(i + 1) + '\n' +
'$D_{KL}[p \ || \ q_{rev}] = ' +
str(np.round(q_rev_loss.numpy(), 4)) + '$',
loc='left')
last_q_rev_loss = q_rev_loss.numpy()
ani = animation.FuncAnimation(fig, animate, frames=num_train_steps)
plt.close()
In [21]:
ani.save('./image/kl_pq.gif', writer='imagemagick', fps=30)

In [22]:
# Plot q and q_rev alongside one another
f, axs = plt.subplots(1, 2, figsize=(15, 7))
axs[0] = plot_density_contours(p.prob, X1, X2,
{'levels':contour_levels,
'cmap':'cividis', 'alpha':0.5}, ax=axs[0])
axs[0] = plot_density_contours(q.prob, X1, X2,
{'levels':contour_levels,
'cmap':'plasma'}, ax=axs[0])
axs[0].set_title('Density contours of $p$ and $q$\n' +
'$D_{KL}[q \ || \ p] = ' + str(np.round(last_q_loss, 4)) + '$',
loc='left')
axs[1] = plot_density_contours(p.prob, X1, X2,
{'levels':contour_levels,
'cmap':'cividis', 'alpha':0.5}, ax=axs[1])
axs[1] = plot_density_contours(q_rev.prob, X1, X2,
{'levels':contour_levels,
'cmap':'plasma'}, ax=axs[1])
axs[1].set_title('Density contours of $p$ and $q_{rev}$\n' +
'$D_{KL}[p \ || \ q_{rev}] = ' + str(np.round(last_q_rev_loss, 4)) + '$',
loc='left')
plt.show()