
 The new Python client library of ServiceX, the novel data delivery system
The new Python client library of ServiceX, the novel data delivery system
KyungEon Choi (UT Austin) for ServiceX team (IRIS-HEP)
PyHEP 2024 (July 1, 2024)
ServiceX
ServiceX is a scalable data extraction, transformation and delivery system deployed in a Kubernetes cluster.
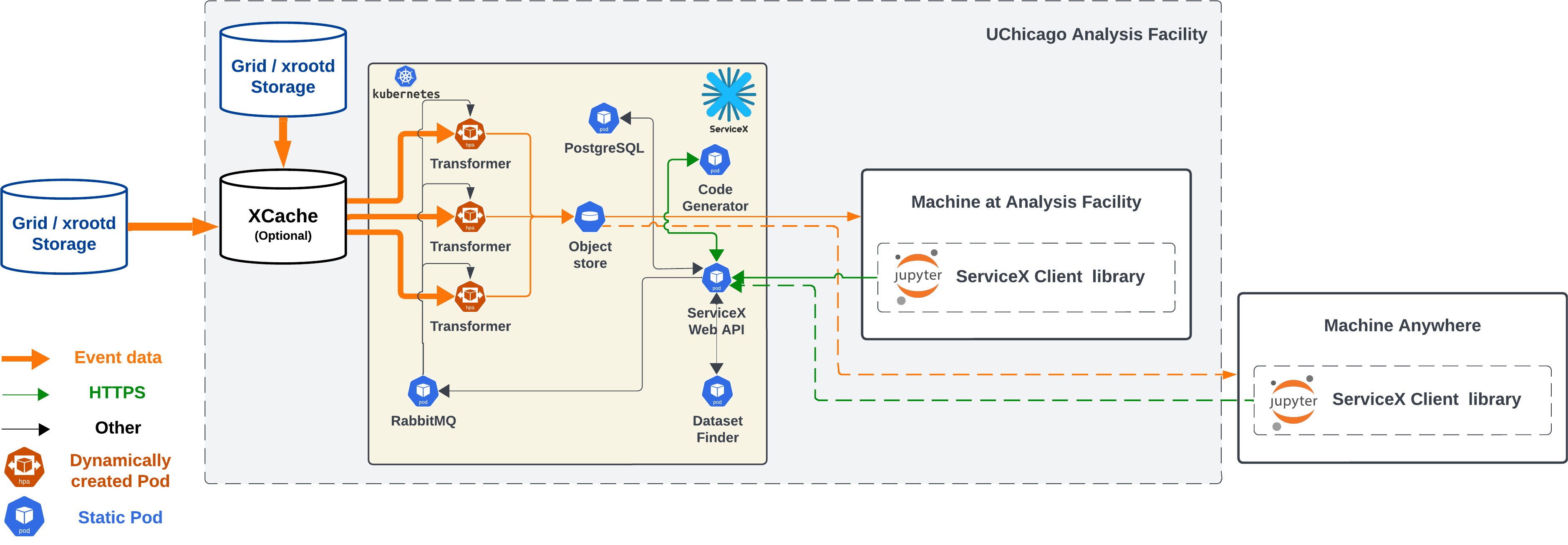
- Event data
- ServiceX delivers from grid or remote XRootD storage to the user. Or more precisely ServiceX writes into an object store (ServiceX internal storage) and users download files or URLs from the object store as soon as available.
- Thickness of arrows reflect the amount of data over a wire. ServiceX is NOT designed to download full data from grids. Transformers effectively reduce data that will be delivered to user based on a query for selection and filtering.
- ServiceX is often co-located with a grid site to maximize network bandwith. XCache is preferable to allow much faster read for frequently accessed datasets.
- Transformer
- ServiceX consists of multiple microservices that are deployed as static K8s pod (always "running" state) but transformers are dynamically created via HPA (Horizontal Pod Scaling)
- A transformer pod runs on a file at a time and number of transformer pods are scaled up and down depending on the number of input files in the dataset and other criteria.
- ServiceX Request
- ServiceX request(s) is(are) made from the SerivceX client libary to ServiceX Web API via HTTP request
- A ServiceX request takes one input dataset (or list of files) and ServiceX is happily scale transformer pods automatically. A dataset with a single file should work but it's much more desirable to utilize HPA.
- Users can make ServiceX request anywhere only with Python ServiceX client library and
servicex.yamlincludes an access token. Thus it's perfectly fine to deliver data to a university cluster or a laptop for small tests.
ServiceX Webpage
- Download a ServiceX configuration file (
servicex.yaml) from the ServiceX website and copy to your home or working directory - NOTE: the ServiceX endpoint
servicex.af.uchicago.eduis limited to the ATLAS users as it provides an access to the ATLAS event data
ServiceX Client library
ServiceX Client library is a python library for users to communicate with ServiceX backend (or server) to make delivery requests and handling of outputs
The most fundamental compenents of a ServiceX request
- Dataset
- Query - describe what a user wants to run in transformers
Design goal of the new ServiceX Client library
- Minimize boilerplates
- Support YAML interface (integration of ServiceX DataBinder)
- Strongly typed (pydantic)
Installation
pip install servicex==3.0.0.alpha.18
In [1]:
# !pip install servicex==3.0.0.alpha.18
!pip list | grep servicex
servicex 3.0.0a18
I have downloaded my ServiceX configuration file (
--> Ready to make a ServiceX request!
servicex.yaml) from the ServiceX webpage and installed servicex package
--> Ready to make a ServiceX request!
First ServiceX request
Let's begin with the basic:Deliver a branch (or column) from a dataset in the grid
In [2]:
import servicex
In [3]:
spec = {
"Sample":[{
"Name": "UprootRaw_PyHEP",
"Dataset": servicex.dataset.Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": servicex.query.UprootRaw({"treename": "nominal", "filter_name": "el_pt"})
}]
}
- One sample named "UprootRaw_PyHEP" is defined in the
specobject. - A Rucio dataset is specified
- Defined a
Query, sent to transformers and run on all files in the given Rucio dataset UprootRawquery takes"treename"to setTTreein flat ROOT ntuples and"filter_name"to select branches in a given tree
Let's deliver my ServiceX request
In [4]:
o = servicex.deliver(spec)
Output()
In [5]:
len(o['UprootRaw_PyHEP'])
Out[5]:
3
Returns a dictionary
In [6]:
print(f"Sample.Name: {o.keys()}\n")
print(f"Fileset: {type(o['UprootRaw_PyHEP'])}\n")
print(f"First file: {(o['UprootRaw_PyHEP'][0])}\n")
Sample.Name: dict_keys(['UprootRaw_PyHEP']) Fileset: <class 'list'> First file: /Users/kc43627/Work/data/servicex_cache/c9a57bae-b2c3-4432-93cd-253763e42ead/root___192.170.240.145__root___fax.mwt2.org_1094__pnfs_uchicago.edu_atlaslocalgroupdisk_rucio_user_mgeyik_a0_3c_user.mgeyik.30183079._000006.out.root
In [7]:
import uproot
with uproot.open(o['UprootRaw_PyHEP'][0]) as f:
column = f['nominal']['el_pt']
column.array()
Out[7]:
[[3.86e+04, 3.6e+04], [3.44e+04, 1.91e+04], [], [5.98e+04, 5.76e+04], [6.84e+04, 2.4e+04], [3.5e+04], [], [], [1.34e+05, 4.36e+04], [], ..., [6.46e+04, 3.66e+04, 2.78e+04], [], [1.74e+04], [], [5.42e+04], [3.81e+04, 1.26e+04], [], [3.53e+04], [6.17e+04]] -------------------------------- type: 11543 * var * float32
Only few lines of a python script brings the data you want from the grid!
Let me go through what kinds of Dataset and Query are supported by ServiceX
Dataset
ServiceX supports Rucio, XRootD, and CERN OpenDataset
In [8]:
servicex.dataset.Rucio.__init__
Out[8]:
<function servicex.dataset_identifier.RucioDatasetIdentifier.__init__(self, dataset: str, num_files: Optional[int] = None)>
In [9]:
servicex.dataset.FileList.__init__
Out[9]:
<function servicex.dataset_identifier.FileListDataset.__init__(self, files: Union[List[str], str])>
In [10]:
servicex.dataset.CERNOpenData.__init__
Out[10]:
<function servicex.dataset_identifier.CERNOpenDataDatasetIdentifier.__init__(self, dataset: int, num_files: Optional[int] = None)>
Query
- Query is a representation of what user wants from input dataset. e.g.
UprootRaw({"treename": "nominal", "filter_name": "el_pt"})- User provided query is translated into a code that runs on transformers
- Query is input data format dependent as a code for flat ROOT ntuple differs from the one for Apache parquet
In [11]:
servicex.query.plugins
Out[11]:
[EntryPoint(name='FuncADL_ATLASr21', value='servicex.func_adl.func_adl_dataset:FuncADLQuery_ATLASr21', group='servicex.query'), EntryPoint(name='FuncADL_ATLASr22', value='servicex.func_adl.func_adl_dataset:FuncADLQuery_ATLASr22', group='servicex.query'), EntryPoint(name='FuncADL_ATLASxAOD', value='servicex.func_adl.func_adl_dataset:FuncADLQuery_ATLASxAOD', group='servicex.query'), EntryPoint(name='FuncADL_CMS', value='servicex.func_adl.func_adl_dataset:FuncADLQuery_CMS', group='servicex.query'), EntryPoint(name='FuncADL_Uproot', value='servicex.func_adl.func_adl_dataset:FuncADLQuery_Uproot', group='servicex.query'), EntryPoint(name='PythonFunction', value='servicex.python_dataset:PythonQuery', group='servicex.query'), EntryPoint(name='UprootRaw', value='servicex.uproot_raw.uproot_raw:UprootRawQuery', group='servicex.query')]
Query classes for ROOT ntuples (via Uproot)
UprootRaw Query
- This is a new query language, essentially calling
uproot.tree.arrays()function - A UprootRaw query can be a dictionary or a list of dictionaries
- There are two types of operations a user can put in a dictionary
- query: contains a
treenamekey - copy: contains a
copy_histogramskey
- query: contains a
query = [
{
'treename': 'reco',
'filter_name': ['/mu.*/', 'runNumber', 'lbn', 'jet_pt_*'],
'cut':'(count_nonzero(jet_pt_NOSYS>40e3, axis=1)>=4)'
},
{
'copy_histograms': ['CutBookkeeper*', '/cflow.*/', 'metadata', 'listOfSystematics']
}
]
- More details on the grammar can be found here
In [12]:
query_UprootRaw = servicex.query.UprootRaw({"treename": "nominal", "filter_name": "el_pt"})
FuncADL_Uproot Query
- Functional Analysis Description Language is a powerful query language that has been supported by ServiceX
- In addition to the basic operations like
Select()for column selection orWhere()for filtering, more sophisticated query can be built - One new addition
FromTree()method to set a tree name in a query - More details can be found at the talk by M. Proffitt at PyHEP 2021
In [13]:
query_FuncADL = servicex.query.FuncADL_Uproot().FromTree('nominal').Select(lambda e: {'el_pt': e['el_pt']})
PythonFunction Query
- Python function can be passed as a query
uproot,awkward,vectorcan be imported (limited by the transformer image)- Primarily experimental purpose and likely to be discontinued
In [14]:
def run_query(input_filenames=None):
import uproot
with uproot.open({input_filenames: "nominal"}) as o:
br = o.arrays("el_pt")
return br
query_PythonFunction = servicex.query.PythonFunction().with_uproot_function(run_query)
All three queries return the same output, ROOT files with selected branch
el_pt_NOSYS!
Multiple samples
- HEP analysis often needs more than one sample
In [15]:
spec_multiple = {
"Sample":[
{
"Name": "UprootRaw_PyHEP",
"Dataset": servicex.dataset.Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": query_UprootRaw
},
{
"Name": "FuncADL_Uproot_PyHEP",
"Dataset": servicex.dataset.Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": query_FuncADL
},
{
"Name": "PythonFunction_PyHEP",
"Dataset": servicex.dataset.Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": query_PythonFunction
}
]
}
Sampleblock is a list of dictionaries, each with aDataset-Querypair- Client library makes one ServiceX request per
Dataset-Querypair - Again, it's preferred to have more files in a request to utilize K8s HPA than having multiple requests for the same query
In [16]:
o_multiple = servicex.deliver(spec_multiple)
Output()
YAML interface
- It's cool to deliver only interested columns from grid storages in a Jupyter notebook, but real analysis often becomes quite messy
- The new client library brings
servicex-databinderand significantly improve user interface to allow a seamless experience with YAML
In [21]:
%%writefile -a config_UprootRaw.yaml
Sample:
- Name: Uproot_UprootRaw_YAML
Dataset: !Rucio user.kchoi.pyhep2024.test_dataset
Query: !UprootRaw |
{"treename":"nominal", "filter_name": "el_pt"}
Writing config_UprootRaw.yaml
Compare with the one in this notebook
In [22]:
from servicex.dataset import Rucio
from servicex.query import UprootRaw
from servicex import deliver
spec = {
"Sample":[{
"Name": "UprootRaw_PyHEP",
"Dataset": Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": UprootRaw({"treename": "nominal", "filter_name": "el_pt"})
}]
}
In [23]:
from servicex import deliver
In [24]:
o_yaml = deliver("config_UprootRaw.yaml")
# o_py = deliver(spec)
Output()
YAML syntax
- The exclamation mark(!) to declare dataset type and query type (see detail on the PyYAML constructor)
- Dataset tags:
!Rucio,!Rucio,!FileList,!CERNOpenData - Query tags:
!UprootRaw,!FuncADL_Uproot,!PythonFunction
- Dataset tags:
- The pipe (
|) after query tag represents the literal operator and allows to properly interpret multi-line string
Optional configurations
Definition:
- &DEF_ggH_input "root://eospublic.cern.ch//eos/opendata/atlas/OutreachDatasets\
/2020-01-22/4lep/MC/mc_345060.ggH125_ZZ4lep.4lep.root"
- &DEF_query1 !PythonFunction |
def run_query(input_filenames=None):
import uproot
with uproot.open({input_filenames:"nominal"}) as o:
br = o.arrays("mu_pt")
return br
- &DEF_query2 !FuncADL_Uproot |
FromTree('mini').Select(lambda e: {'lep_pt': e['lep_pt']}).Where(lambda e: e['lep_pt'] > 1000)
General:
OutputFormat: parquet
Delivery: SignedURLs
Sample:
- Name: ttH
Dataset: !Rucio user.kchoi.fcnc_tHq_ML.ttH.v11
Query: *DEF_query1
NFiles: 5
# IgnoreLocalCache: False
- Name: ttZ
Dataset: !Rucio user.kchoi.fcnc_tHq_ML.ttZ.v11
Query: *DEF_query1
NFiles: 3
- Name: ggH
Dataset: !FileList *DEF_ggH_input
Query: *DEF_query2
Failed transformation
In [25]:
spec_typo = {
"Sample":[{
"Name": "UprootRaw_PyHEP",
"Dataset": Rucio("user.kchoi.pyhep2024.test_dataset"),
"Query": UprootRaw({"treename": "nominal", "filter_name": "el_pta"})
}]
}
In [26]:
o = deliver(spec_typo)
Output()
[07/01/24 00:13:17] WARNING Transform "UprootRaw_PyHEP" completed with failures: 3/3 files query_core.py:215 failed
WARNING More information of 'UprootRaw_PyHEP' HERE query_core.py:226
Future plans¶
Client library
- Improve robustness: progress bar (transform status/object store access) and local caching
- Readthedoc of the new ServiceX cilent library is under construction! https://servicex-frontend.readthedocs.io/en/latest/index.html
ServiceX
- Improve stability and robustness of ServiceX especially what we learned during 200Gbps challenge (hundreds of ServiceX requests on hundreds of TB datasets)
- Server-side caching
- Other ServiceX transformers: ATLAS TopCPToolkit transformer, column-join transformer
In [ ]: