Structural distinction using pyiron and pyscal¶
In this tutorial, we will read in MD snapshots using pyiron. We will then use the pyscal module to indentify the structure in which the MD configuration is in.
Import necessary modules
import os
import pyiron
from ase.io import read
import pyscal.core as pc
import matplotlib.pyplot as plt
Create a pyiron project
pr = pyiron.Project('ADIS')
Check the job table
pr.job_table()
| id | status | chemicalformula | job | subjob | projectpath | project | timestart | timestop | totalcputime | computer | hamilton | hamversion | parentid | masterid | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 2 | finished | H8192 | Nucleus | /Nucleus | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:34.985406 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 1 | 4 | finished | H3980 | Si_1400_20 | /Si_1400_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:40.585864 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 2 | 5 | finished | H1529 | S5_s00_n130_r1024_720_5 | /S5_s00_n130_r1024_720_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:43.147042 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 3 | 6 | finished | H1980 | Fe_1050_20 | /Fe_1050_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:47.255863 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 4 | 7 | finished | H1980 | Fe_1350_20 | /Fe_1350_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:53.247038 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 5 | 8 | finished | H3995 | Si_1000_5 | /Si_1000_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:11:59.993315 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 6 | 9 | finished | H1980 | Cu_720_20 | /Cu_720_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:06.132659 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 7 | 10 | finished | H2000 | Cu_400_0 | /Cu_400_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:11.172906 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 8 | 11 | finished | H4000 | Mg_900_0 | /Mg_900_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:18.633493 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 9 | 12 | finished | H4000 | Mg_500_0 | /Mg_500_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:24.458570 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 10 | 13 | finished | H1534 | S5_s00_n130_r1024_720_0 | /S5_s00_n130_r1024_720_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:26.942533 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 11 | 15 | finished | H3995 | Mg_700_5 | /Mg_700_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:33.466119 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 12 | 16 | finished | H3980 | Si_1000_20 | /Si_1000_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:40.492150 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 13 | 17 | finished | H1554 | S5_s00_n130_r1024_560_5 | /S5_s00_n130_r1024_560_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:43.054725 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 14 | 18 | finished | H1584 | S5_s00_n130_r1024_400_0 | /S5_s00_n130_r1024_400_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:44.725043 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 15 | 19 | finished | H1980 | Cu_400_20 | /Cu_400_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:49.268366 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 16 | 20 | finished | H1995 | Fe_1350_5 | /Fe_1350_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:12:54.623718 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 17 | 21 | finished | H3995 | Mg_1000_5 | /Mg_1000_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:01.055022 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 18 | 22 | finished | H4000 | Si_1400_0 | /Si_1400_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:09.023513 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 19 | 23 | finished | H2000 | Fe_1350_0 | /Fe_1350_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:14.466088 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 20 | 24 | finished | H4000 | Mg_700_0 | /Mg_700_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:21.071463 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 21 | 26 | finished | H1995 | Fe_1050_5 | /Fe_1050_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:26.326193 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 22 | 27 | finished | H3980 | Mg_1000_20 | /Mg_1000_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:34.325798 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 23 | 28 | finished | H2000 | Cu_560_0 | /Cu_560_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:40.855803 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 24 | 29 | finished | H2000 | Fe_750_0 | /Fe_750_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:48.369252 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 25 | 30 | finished | H1995 | Cu_400_5 | /Cu_400_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:55.268210 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 26 | 31 | finished | H1579 | S5_s00_n130_r1024_400_5 | /S5_s00_n130_r1024_400_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:13:57.530915 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 27 | 32 | finished | H1980 | Fe_750_20 | /Fe_750_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:01.649228 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 28 | 33 | finished | H1980 | Cu_560_20 | /Cu_560_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:06.227415 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 29 | 34 | finished | H2000 | Cu_720_0 | /Cu_720_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:10.660001 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 30 | 35 | finished | H3980 | Mg_500_20 | /Mg_500_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:15.234711 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 31 | 36 | finished | H3980 | Mg_700_20 | /Mg_700_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:19.744690 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 32 | 38 | finished | H3995 | Mg_500_5 | /Mg_500_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:25.832014 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 33 | 39 | finished | H500 | Liquid | /Liquid | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:28.167739 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 34 | 40 | finished | H1995 | Cu_560_5 | /Cu_560_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:34.625711 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 35 | 41 | finished | H1510 | S5_s00_n130_r1024_720_20 | /S5_s00_n130_r1024_720_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:37.464296 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 36 | 42 | finished | H3980 | Mg_900_20 | /Mg_900_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:45.426443 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 37 | 43 | finished | H2000 | Fe_1050_0 | /Fe_1050_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:52.439612 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 38 | 44 | finished | H1559 | S5_s00_n130_r1024_560_0 | /S5_s00_n130_r1024_560_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:14:55.155021 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 39 | 45 | finished | H4000 | Mg_1000_0 | /Mg_1000_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:02.737287 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 40 | 46 | finished | H1534 | S5_s00_n130_r1024_560_20 | /S5_s00_n130_r1024_560_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:06.033125 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 41 | 47 | finished | H3995 | Si_1400_5 | /Si_1400_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:13.332531 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 42 | 48 | finished | H1995 | Cu_720_5 | /Cu_720_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:21.831887 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 43 | 49 | finished | H1995 | Fe_750_5 | /Fe_750_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:29.351879 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 44 | 50 | finished | H4000 | Si_1000_0 | /Si_1000_0 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:37.769224 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 45 | 51 | finished | H1559 | S5_s00_n130_r1024_400_20 | /S5_s00_n130_r1024_400_20 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:39.951420 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
| 46 | 53 | finished | H3995 | Mg_900_5 | /Mg_900_5 | /home/jovyan/ | ADIS/ | 2020-11-05 21:15:46.458817 | None | None | pyiron@jupyter-pyiron-2dadis-2dworkshop-2d2020-2d3piumz95#1 | AtomisticGenericJob | 0.2.0 | None | None |
We can try an FCC structure, Copper at 400 K
job = pr.load("Cu_400_0")
Get the structure from the job
Cu400 = job.get_structure()
Common Neighbor Analysis¶
This is a simple FCC structure, so we can an adaptive common neighbor analysis method to find the structure. CNA uses three indices for each pair of atoms, let's say $i$ and $j$:
- The number of common neighbors shared by atoms $i$ and $j$
- The number of bonds among these common neighbors
- The longest chain of bonds connecting these neighbors
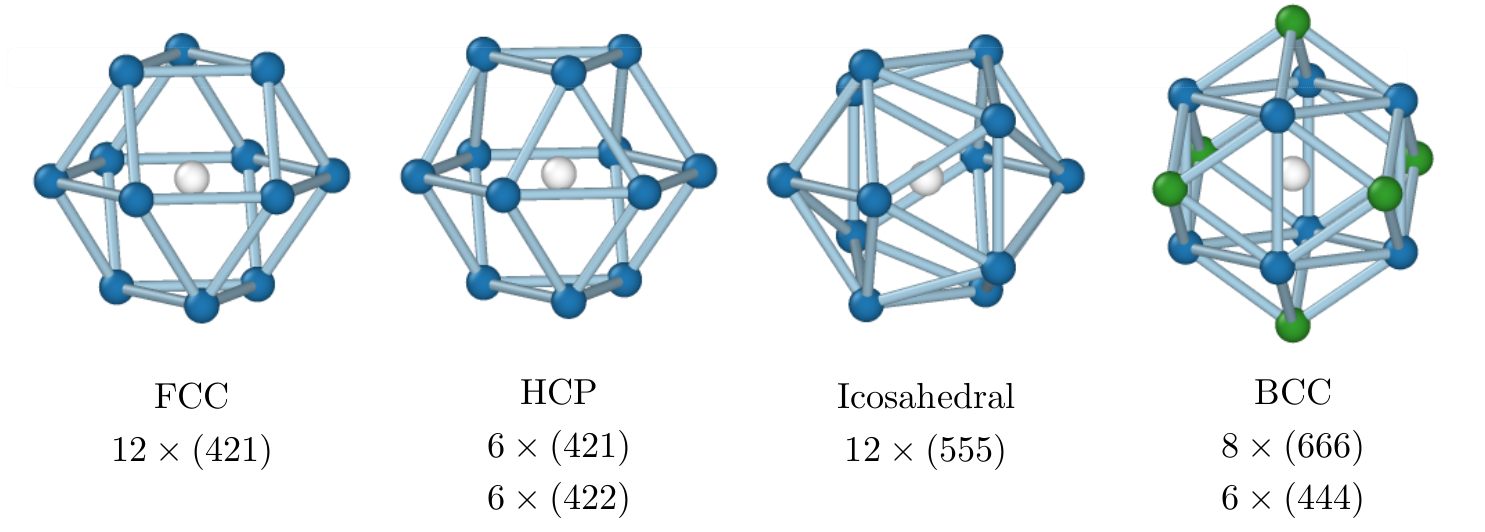
Image source: arXiv:2003.08879
For the analysis, we will use pyscal as it is implemented within pyiron
| pyscal | |
| pyscal is a python module for the calculation of local atomic structural environments including Steinhardt's bond orientational order parameters during post-processing of atomistic simulation data. The core functionality of pyscal is written in C++ with python wrappers using pybind11 which allows for fast calculations with possibilities for easy expansion in python. |
 |
from pyiron.atomistics.structure.pyscal import analyse_cna_adaptive
analyse_cna_adaptive(Cu400)
[17, 1983, 0, 0, 0]
The numbers are in the order : Others, fcc, hcp, bcc, icosahedral. From the numbers, it is clear that the system is completely fcc structured.
Let's try for a different structure. Cu at 720 K.
job = pr.load("Cu_720_0")
Cu720 = job.get_structure()
analyse_cna_adaptive(Cu720)
[133, 1867, 0, 0, 0]
When we follow the same procedure, theres only 1835 atoms. 165 atoms are now identified as Others. Structural identification becomes increasingly difficult at higher temperatures.
Steinhardt's parameters¶
Steinhardt's bond orientational order parameters are a set of parameters based on spherical harmonics to explore the local atomic environment. These parameters have been used extensively for various uses such as distinction of crystal structures, identification of solid and liquid atoms and identification of defects.
These parameters, which are rotationally and translationally invariant are defined by,
$$ q_l (i) = \Big( \frac{4\pi}{2l+1} \sum_{m=-l}^l | q_{lm}(i) |^2 \Big )^{\frac{1}{2}} $$
where,
$$ q_{lm} (i) = \frac{1}{N(i)} \sum_{j=1}^{N(i)} Y_{lm}(\pmb{r}_{ij}) $$
in which $Y_{lm}$ are the spherical harmonics and $N(i)$ is the number of neighbours of particle $i$, $\pmb{r}_{ij}$ is the vector connecting particles $i$ and $j$, and $l$ and $m$ are both intergers with $m \in [-l,+l]$. Various parameters have found specific uses, such as $q_2$ and $q_6$ for identification of crystallinity, $q_6$ for identification of solidity, and $q_4$ and $q_6$ for distinction of crystal structures.
First a function to calculate Steinhardt's parameters
from pyiron.atomistics.structure.pyscal import get_steinhardt_parameter_job
We will take the basic lattices - bcc, fcc and hcp and see how the Steinhardt's parameters look
bccq, _ = get_steinhardt_parameter_job(pr.load("Fe_750_0"), cutoff=3.3, clustering=False)
fccq, _ = get_steinhardt_parameter_job(pr.load("Cu_400_0"), clustering=False)
hcpq, _ = get_steinhardt_parameter_job(pr.load("Mg_500_0"), clustering=False)
Lets plot the values
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
plt.xlabel(r"$q_4$")
plt.ylabel(r"$q_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79e93ea610>
The distributions look well separated. Lets try at a higher temperature.
bccq, _ = get_steinhardt_parameter_job(pr.load("Fe_1350_0"), cutoff=3.3, clustering=False)
fccq, _ = get_steinhardt_parameter_job(pr.load("Cu_720_0"), clustering=False)
hcpq, _ = get_steinhardt_parameter_job(pr.load("Mg_1000_0"), clustering=False)
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
plt.xlabel(r"$q_4$")
plt.ylabel(r"$q_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79e92c08d0>
It gets worse due to thermal fluctuations, but we can averaged the values over the neighbors
bccq, _ = get_steinhardt_parameter_job(pr.load("Fe_1350_0"), cutoff=3.3, averaged=True, clustering=False)
fccq, _ = get_steinhardt_parameter_job(pr.load("Cu_720_0"), averaged=True, clustering=False)
hcpq, _ = get_steinhardt_parameter_job(pr.load("Mg_1000_0"), averaged=True, clustering=False)
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
plt.xlabel(r"$\bar{q}_4$")
plt.ylabel(r"$\bar{q}_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79f2e7b450>
We can also try adding a diamond structure to the mix
diaq, _ = get_steinhardt_parameter_job(pr.load("Si_1400_0"), averaged=True, clustering=False)
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
plt.scatter(diaq[0], diaq[1], alpha=0.05, label="dia", color="#00838F")
plt.xlabel(r"$\bar{q}_4$")
plt.ylabel(r"$\bar{q}_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79e9125050>
Add a liquid structure
lqdq, _ = get_steinhardt_parameter_job(pr.load("Liquid"), averaged=True, clustering=False)
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
plt.scatter(diaq[0], diaq[1], alpha=0.05, label="dia", color="#00838F")
plt.scatter(lqdq[0], lqdq[1], alpha=0.20, label="lqd", color="#1E88E5")
plt.xlabel(r"$\bar{q}_4$")
plt.ylabel(r"$\bar{q}_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79e83d5b50>
bccq, _ = get_steinhardt_parameter_job(pr.load("Fe_1350_0"), q=(8, 12), cutoff=3.3, averaged=True, clustering=False)
fccq, _ = get_steinhardt_parameter_job(pr.load("Cu_720_0"), q=(8, 12),averaged=True, clustering=False)
hcpq, _ = get_steinhardt_parameter_job(pr.load("Mg_1000_0"), q=(8, 12),averaged=True, clustering=False)
plt.scatter(bccq[0], bccq[1], alpha=0.05, label="bcc", color="#B71C1C")
plt.scatter(fccq[0], fccq[1], alpha=0.05, label="fcc", color="#F57F17")
plt.scatter(hcpq[0], hcpq[1], alpha=0.05, label="hcp", color="#1B5E20")
#plt.scatter(diaq[0], diaq[1], alpha=0.05, label="dia", color="#00838F")
#plt.scatter(lqdq[0], lqdq[1], alpha=0.20, label="lqd", color="#1E88E5")
plt.xlabel(r"$\bar{q}_4$")
plt.ylabel(r"$\bar{q}_6$")
plt.legend()
<matplotlib.legend.Legend at 0x7f79e92dd050>
Get all q values
bccq, _ = get_steinhardt_parameter_job(pr.load("Fe_1350_5"), q=range(2, 13), cutoff=3.3, averaged=True, clustering=False)
fccq, _ = get_steinhardt_parameter_job(pr.load("Cu_720_0"), q=range(2, 13),averaged=True, clustering=False)
hcpq, _ = get_steinhardt_parameter_job(pr.load("Mg_1000_0"), q=range(2, 13),averaged=True, clustering=False)
Import clustering
import sklearn.cluster
import numpy as np
c1 = sklearn.cluster.k_means(np.hstack([bccq[:2], fccq[:2], hcpq[:2]]).T, 3)
wrong_bcc = np.sum(c1[1][:2000] != np.median(c1[1][:2000]))
wrong_fcc = np.sum(c1[1][2000:4000] != np.median(c1[1][2000:4000]))
wrong_hcp = np.sum(c1[1][4000:8000] != np.median(c1[1][4000:8000]))
acc = 1 - (wrong_bcc +wrong_fcc + wrong_hcp)/8000
acc
0.73925
Get accuracy measure
for i in range(2, 13):
c1 = sklearn.cluster.k_means(np.hstack([bccq[:i], fccq[:i], hcpq[:i]]).T, 3)
wrong_bcc = np.sum(c1[1][:2000] != np.median(c1[1][:2000]))
wrong_fcc = np.sum(c1[1][2000:4000] != np.median(c1[1][2000:4000]))
wrong_hcp = np.sum(c1[1][4000:8000] != np.median(c1[1][4000:8000]))
acc = 1 - (wrong_bcc +wrong_fcc + wrong_hcp)/8000
print(acc, c1[2]/i)
0.73975 0.2585592276191804 0.987625 0.6129001784949375 0.993125 0.8039044257798553 0.998625 1.089862946377139 0.998375 1.1567243159208072 0.99875 1.2987743228155133 0.99875 1.2215957539637534 0.99875 1.2861025616220654 0.99875 1.3203286907380984 0.998625 2.1627207520443092 0.998625 1.9824940227072834
Check combination of 4 and 6
c1 = sklearn.cluster.k_means(np.hstack([np.array(bccq)[[2,4]], np.array(fccq)[[2,4]], np.array(hcpq)[[2,4]]]).T, 3)
wrong_bcc = np.sum(c1[1][:2000] != np.median(c1[1][:2000]))
wrong_fcc = np.sum(c1[1][2000:4000] != np.median(c1[1][2000:4000]))
wrong_hcp = np.sum(c1[1][4000:8000] != np.median(c1[1][4000:8000]))
acc = 1 - (wrong_bcc +wrong_fcc + wrong_hcp)/8000
print(acc, c1[2]/2)
0.99875 1.6522672168101433
Check combination of 8 and 12
c1 = sklearn.cluster.k_means(np.hstack([np.array(bccq)[[6,10]], np.array(fccq)[[6,10]], np.array(hcpq)[[6,10]]]).T, 3)
wrong_bcc = np.sum(c1[1][:2000] != np.median(c1[1][:2000]))
wrong_fcc = np.sum(c1[1][2000:4000] != np.median(c1[1][2000:4000]))
wrong_hcp = np.sum(c1[1][4000:8000] != np.median(c1[1][4000:8000]))
acc = 1 - (wrong_bcc +wrong_fcc + wrong_hcp)/8000
print(acc, c1[2]/2)
0.992875 6.262579674544909
Check combination of even qs
c1 = sklearn.cluster.k_means(np.hstack([np.array(bccq)[[2, 4, 6,10]], np.array(fccq)[[2, 4, 6,10]], np.array(hcpq)[[2, 4, 6,10]]]).T, 3)
wrong_bcc = np.sum(c1[1][:2000] != np.median(c1[1][:2000]))
wrong_fcc = np.sum(c1[1][2000:4000] != np.median(c1[1][2000:4000]))
wrong_hcp = np.sum(c1[1][4000:8000] != np.median(c1[1][4000:8000]))
acc = 1 - (wrong_bcc +wrong_fcc + wrong_hcp)/8000
print(acc, c1[2]/5)
0.99875 3.208414883331198